Database Schema
ERDs and table documentation for SDM and SPP databases
ERDs and table documentation for SDM and SPP databases
2026-02-13 13:43:45
---
title: "Database Schema"
subtitle: "ERDs and table documentation for SDM and SPP databases"
format:
html:
mermaid-format: png
editor_options:
chunk_output_type: console
---
```{r}
#| label: setup
#| include: false
librarian::shelf(
DBI,
duckdb,
DT,
dplyr,
glue,
purrr,
tibble,
quiet = T
)
# paths ----
is_server <- Sys.info()[["sysname"]] == "Linux"
dir_data <- ifelse(is_server, "/share/data", "~/My Drive/projects/msens/data")
dir_big <- ifelse(is_server, "/share/data/big", "~/_big/msens/derived")
dir_big_v <- glue("{dir_big}/v3")
# connections (read-only) ----
con_sdm <- dbConnect(duckdb(
dbdir = glue("{dir_big_v}/sdm.duckdb"),
read_only = TRUE
))
con_spp <- dbConnect(duckdb(
dbdir = glue("{dir_big}/spp.duckdb"),
read_only = TRUE
))
# expected schema ----
# update these lists when tables or columns are added/removed
sdm_expected <- list(
cell = c(
"cell_id",
"depth_mean",
"depth_min",
"depth_max",
"oxy_b_mean",
"oxy_mean",
"prim_prod_mean",
"ice_con_ann",
"salinity_b_mean",
"salinity_mean",
"sbt_an_mean",
"sst_an_mean",
"fao_area_m",
"area_km2",
"width_km2"
),
cell_metric = c("cell_id", "metric_seq", "value"),
dataset = c(
"ds_key",
"name_short",
"name_original",
"description",
"citation",
"source_broad",
"source_detail",
"regions",
"response_type",
"taxa_groups",
"year_pub",
"date_obs_beg",
"date_obs_end",
"date_env_beg",
"date_env_end",
"link_info",
"link_download",
"link_metadata",
"links_other",
"spatial_res_deg",
"temporal_res",
"date_created",
"name_display",
"value_info",
"is_mask",
"sort_order"
),
listing = c(
"spp_id",
"worms_id",
"botw_id",
"extrisk_code",
"er_score",
"is_mmpa",
"is_mbta",
"is_bcc"
),
metric = c("metric_seq", "metric_key", "description", "date_created"),
model = c(
"mdl_seq",
"ds_key",
"taxa",
"time_period",
"region",
"mdl_type",
"description",
"date_created"
),
model_cell = c("mdl_seq", "cell_id", "value"),
species = c(
"sp_seq",
"ds_key",
"taxa",
"sp_key",
"worms_id",
"gbif_id",
"itis_id",
"scientific_name_dataset",
"common_name_dataset",
"scientific_name_accepted",
"common_name_accepted",
"iucn_id",
"redlist_code",
"redlist_year",
"sp_cat",
"worms_is_marine",
"worms_is_extinct",
"botw_id"
),
taxon = c(
"taxon_id",
"taxon_authority",
"n_ds",
"ms_merge",
"sp_cat",
"mdl_seq",
"scientific_name",
"common_name",
"worms_id",
"redlist_code",
"extrisk_code",
"er_score",
"is_mmpa",
"is_mbta",
"is_bcc",
"worms_is_marine",
"worms_is_extinct",
"esa_code",
"esa_source",
"is_ok"
),
taxon_model = c("taxon_id", "ds_key", "mdl_seq"),
zone = c("zone_seq", "tbl", "fld", "value", "date_created"),
zone_cell = c("zone_seq", "cell_id", "pct_covered"),
zone_metric = c("zone_seq", "metric_seq", "value"),
zone_taxon = c(
"zone_tbl",
"zone_fld",
"zone_value",
"mdl_seq",
"sp_cat",
"sp_common",
"sp_scientific",
"taxon_id",
"taxon_authority",
"rl_code",
"er_score",
"area_km2",
"avg_suit",
"suit_rl",
"suit_rl_area",
"cat_suit_rl_area",
"pct_cat"
)
)
spp_expected <- list(
botw = c(
"taxonID",
"scientificName",
"redlist_code",
"kingdom",
"phylum",
"class",
"order",
"family",
"family_common_name",
"subfamily",
"tribe",
"taxonomic_authority",
"taxonomic_sources",
"acceptedNameUsage",
"acceptedNameUsageID"
),
botw_vernacular = c("taxonID", "vernacularName", "isPreferredName"),
gbif = c(
"taxonID",
"datasetID",
"parentNameUsageID",
"acceptedNameUsageID",
"originalNameUsageID",
"scientificName",
"scientificNameAuthorship",
"canonicalName",
"genericName",
"specificEpithet",
"infraspecificEpithet",
"taxonRank",
"nameAccordingTo",
"namePublishedIn",
"taxonomicStatus",
"nomenclaturalStatus",
"taxonRemarks",
"kingdom",
"phylum",
"class",
"order",
"family",
"genus"
),
gbif_vernacular = c(
"taxonID",
"vernacularName",
"language",
"country",
"countryCode",
"sex",
"lifeStage",
"source"
),
itis = c(
"taxonID",
"parentNameUsageID",
"acceptedNameUsageID",
"scientificName",
"scientificNameAuthorship",
"taxonRank",
"taxonomicStatus",
"nomenclaturalStatus",
"completeness",
"namePublishedIn",
"namePublishedInYear"
),
itis_vernacular = c("taxonID", "vernacularName", "language"),
iucn_redlist = c(
"taxon_scientific_name",
"sis_taxon_id",
"year_published",
"latest",
"possibly_extinct",
"possibly_extinct_in_the_wild",
"url",
"red_list_category_code",
"assessment_id",
"code",
"code_type",
"scopes_description",
"scopes_code",
"red_list_category_code_gom",
"year_published_gom",
"assessment_id_gom"
),
iucn_vernacular = c(
"taxonID",
"isPreferredName",
"language",
"vernacularName"
),
worms = c(
"taxonID",
"scientificNameID",
"acceptedNameUsageID",
"parentNameUsageID",
"namePublishedInID",
"scientificName",
"acceptedNameUsage",
"parentNameUsage",
"namePublishedIn",
"namePublishedInYear",
"kingdom",
"phylum",
"class",
"order",
"family",
"genus",
"subgenus",
"specificEpithet",
"infraspecificEpithet",
"taxonRank",
"scientificNameAuthorship",
"nomenclaturalCode",
"taxonomicStatus",
"nomenclaturalStatus",
"modified",
"bibliographicCitation",
"references",
"license",
"rightsHolder",
"datasetName",
"institutionCode",
"datasetID",
"isMarine",
"isFreshwater",
"isTerrestrial",
"isExtinct",
"isBrackish"
),
worms_vernacular = c(
"taxonID",
"vernacularName",
"source",
"language",
"isPreferredName"
)
)
# helper: check schema changes ----
check_schema <- function(con, expected, db_label) {
actual_cols <- dbGetQuery(
con,
"
SELECT table_name, column_name
FROM information_schema.columns
WHERE table_schema = 'main'
ORDER BY table_name, ordinal_position"
)
actual <- split(actual_cols$column_name, actual_cols$table_name)
msgs <- character()
# check for new/missing tables
new_tables <- setdiff(names(actual), names(expected))
missing_tables <- setdiff(names(expected), names(actual))
if (length(new_tables) > 0) {
msgs <- c(
msgs,
glue("**New tables:** {paste(new_tables, collapse = ', ')}")
)
}
if (length(missing_tables) > 0) {
msgs <- c(
msgs,
glue("**Missing tables:** {paste(missing_tables, collapse = ', ')}")
)
}
# check columns for tables present in both
shared <- intersect(names(actual), names(expected))
for (tbl in shared) {
new_cols <- setdiff(actual[[tbl]], expected[[tbl]])
missing_cols <- setdiff(expected[[tbl]], actual[[tbl]])
if (length(new_cols) > 0) {
msgs <- c(
msgs,
glue(
"**`{tbl}`** new columns: {paste(new_cols, collapse = ', ')}"
)
)
}
if (length(missing_cols) > 0) {
msgs <- c(
msgs,
glue(
"**`{tbl}`** missing columns: {paste(missing_cols, collapse = ', ')}"
)
)
}
}
if (length(msgs) > 0) {
cat("::: {.callout-warning}\n")
cat(glue("## {db_label} schema has changed\n\n"))
cat("Update the ERD and expected schema in this file.\n\n")
cat(paste("-", msgs, collapse = "\n"), "\n")
cat(":::\n\n")
}
}
# helper: infer column description ----
infer_desc <- function(col) {
known <- c(
cell_id = "grid cell identifier",
ds_key = "dataset key",
mdl_seq = "model sequence id (auto-increment)",
mdl_type = "model type",
metric_seq = "metric sequence id (auto-increment)",
metric_key = "metric key identifier",
zone_seq = "zone sequence id (auto-increment)",
taxon_id = "taxon identifier",
taxon_authority = "taxonomic authority source",
sp_seq = "species sequence id",
sp_key = "species key in source dataset",
sp_cat = "species category",
spp_id = "species identifier",
worms_id = "WoRMS taxon id",
gbif_id = "GBIF taxon id",
itis_id = "ITIS taxon id",
iucn_id = "IUCN Red List id",
botw_id = "Birds of the World id",
taxonID = "taxon identifier",
sis_taxon_id = "IUCN SIS taxon id",
scientificName = "scientific name",
scientific_name = "scientific name",
scientific_name_dataset = "scientific name from source dataset",
scientific_name_accepted = "accepted scientific name",
common_name = "common name",
common_name_dataset = "common name from source dataset",
common_name_accepted = "accepted common name",
vernacularName = "common/vernacular name",
taxonRank = "taxonomic rank",
taxonomicStatus = "taxonomic status",
nomenclaturalStatus = "nomenclatural status",
acceptedNameUsageID = "accepted name usage id",
acceptedNameUsage = "accepted name usage",
parentNameUsageID = "parent name usage id",
parentNameUsage = "parent name usage",
canonicalName = "canonical name (without authorship)",
scientificNameAuthorship = "scientific name authorship",
redlist_code = "IUCN Red List category code",
red_list_category_code = "IUCN Red List category code",
extrisk_code = "extinction risk code",
er_score = "extinction risk score",
depth_mean = "mean depth (m)",
depth_min = "minimum depth (m)",
depth_max = "maximum depth (m)",
area_km2 = "area in km²",
pct_covered = "percent of cell covered by zone",
value = "numeric value",
description = "description text",
citation = "citation text",
name_short = "short display name",
name_original = "original name from source",
name_display = "display name",
source_broad = "broad source category",
source_detail = "detailed source",
response_type = "model response type",
taxa_groups = "taxonomic groups covered",
taxa = "taxon name",
time_period = "time period",
region = "geographic region",
tbl = "source table name",
fld = "source field name",
n_ds = "number of datasets",
ms_merge = "merge status flag",
isPreferredName = "whether this is the preferred name",
isMarine = "is marine species",
isFreshwater = "is freshwater species",
isTerrestrial = "is terrestrial species",
isExtinct = "is extinct",
isBrackish = "is brackish water species",
is_mmpa = "protected under MMPA",
is_mbta = "protected under MBTA",
is_bcc = "bird of conservation concern",
is_mask = "whether dataset is a mask layer",
is_ok = "passes quality checks",
worms_is_marine = "classified as marine in WoRMS",
worms_is_extinct = "classified as extinct in WoRMS"
)
if (col %in% names(known)) {
return(known[[col]])
}
# suffix/prefix patterns
if (grepl("_id$", col)) {
return("identifier")
}
if (grepl("_key$", col)) {
return("key")
}
if (grepl("_seq$", col)) {
return("sequence id")
}
if (grepl("_mean$", col)) {
return("mean value")
}
if (grepl("_min$", col)) {
return("minimum value")
}
if (grepl("_max$", col)) {
return("maximum value")
}
if (grepl("_ann$", col)) {
return("annual value")
}
if (grepl("^date_", col)) {
return("date")
}
if (grepl("^is_", col)) {
return("boolean flag")
}
if (grepl("^n_", col)) {
return("count")
}
if (grepl("_code$", col)) {
return("code")
}
if (grepl("^link", col)) {
return("URL")
}
if (grepl("_year$", col)) {
return("year")
}
if (grepl("_name", col)) {
return("name")
}
""
}
# helper: table info as DT ----
tbl_info <- function(con, tbl_name) {
# column metadata
cols <- dbGetQuery(
con,
glue(
"
SELECT column_name, data_type
FROM information_schema.columns
WHERE table_schema = 'main' AND table_name = '{tbl_name}'
ORDER BY ordinal_position"
)
)
n_rows <- dbGetQuery(
con,
glue(
'SELECT COUNT(*) as n FROM "{tbl_name}"'
)
)$n
# build stats per column
stats <- map_dfr(seq_len(nrow(cols)), function(i) {
col <- cols$column_name[i]
dtype <- cols$data_type[i]
col_q <- paste0('"', col, '"')
is_float <- grepl("DOUBLE|FLOAT|DECIMAL|NUMERIC", dtype, ignore.case = TRUE)
# range for numeric / date columns
range_str <- NA_character_
if (grepl(
"INT|DOUBLE|FLOAT|DECIMAL|NUMERIC|DATE|TIMESTAMP",
dtype, ignore.case = TRUE
)) {
min_ex <- if (is_float) glue("ROUND(MIN({col_q}), 4)") else glue("MIN({col_q})")
max_ex <- if (is_float) glue("ROUND(MAX({col_q}), 4)") else glue("MAX({col_q})")
rng <- tryCatch(
dbGetQuery(con, glue(
'SELECT {min_ex}::VARCHAR AS mn, {max_ex}::VARCHAR AS mx
FROM "{tbl_name}"')),
error = function(e) NULL)
if (!is.null(rng) && !is.na(rng$mn))
range_str <- glue("[{rng$mn}, {rng$mx}]")
}
# top 5 most frequent values (sample for very large tables)
sample_clause <- if (n_rows > 5e6) "USING SAMPLE 1000000" else ""
val_ex <- if (is_float) glue("ROUND({col_q}, 4)") else col_q
top5 <- tryCatch(
dbGetQuery(con, glue(
'SELECT {val_ex}::VARCHAR AS val, COUNT(*) AS n
FROM "{tbl_name}" {sample_clause}
GROUP BY {val_ex}
ORDER BY n DESC
LIMIT 5')),
error = function(e) NULL)
top5_str <- NA_character_
if (!is.null(top5) && nrow(top5) > 0) {
top5$val <- ifelse(
nchar(top5$val) > 40,
paste0(substr(top5$val, 1, 37), "..."),
top5$val)
top5_str <- paste(
glue("{top5$val} ({format(top5$n, big.mark=',')})"),
collapse = "; ")
}
tibble(
column = col,
type = dtype,
description = infer_desc(col),
range = range_str,
top_values = top5_str
)
})
datatable(
stats,
caption = glue("{tbl_name} \u2014 {format(n_rows, big.mark = ',')} rows"),
options = list(
scrollX = TRUE,
pageLength = 50,
dom = "ft"
),
rownames = FALSE,
escape = FALSE
)
}
```
# SDM — Species Distribution Model
```{r}
#| label: sdm-changes
#| echo: false
#| output: asis
check_schema(con_sdm, sdm_expected, "SDM")
```
## Schema
```{mermaid}
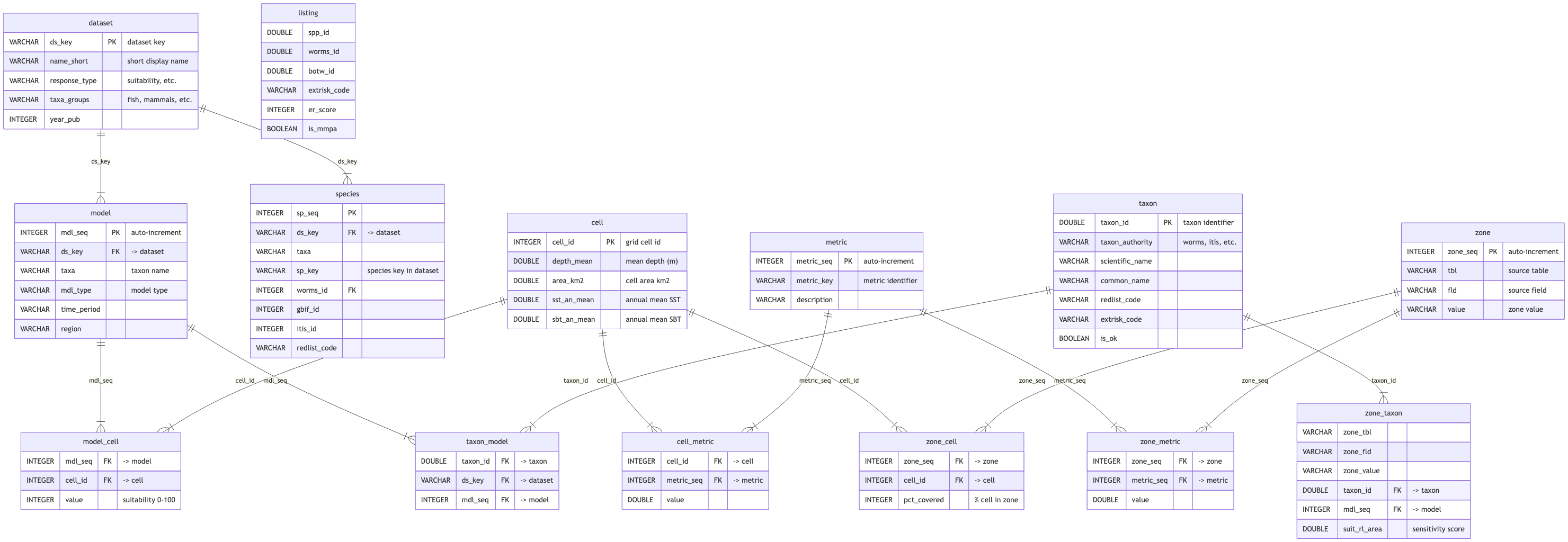
%%| label: fig-sdm-erd
%%| fig-cap: "SDM entity relationship diagram (click to zoom)"
%%| fig-width: 12
erDiagram
dataset {
VARCHAR ds_key PK "dataset key"
VARCHAR name_short "short display name"
VARCHAR response_type "suitability, etc."
VARCHAR taxa_groups "fish, mammals, etc."
INTEGER year_pub
}
model {
INTEGER mdl_seq PK "auto-increment"
VARCHAR ds_key FK "-> dataset"
VARCHAR taxa "taxon name"
VARCHAR mdl_type "model type"
VARCHAR time_period
VARCHAR region
}
species {
INTEGER sp_seq PK
VARCHAR ds_key FK "-> dataset"
VARCHAR taxa
VARCHAR sp_key "species key in dataset"
INTEGER worms_id FK
INTEGER gbif_id
INTEGER itis_id
VARCHAR redlist_code
}
cell {
INTEGER cell_id PK "grid cell id"
DOUBLE depth_mean "mean depth (m)"
DOUBLE area_km2 "cell area km2"
DOUBLE sst_an_mean "annual mean SST"
DOUBLE sbt_an_mean "annual mean SBT"
}
model_cell {
INTEGER mdl_seq FK "-> model"
INTEGER cell_id FK "-> cell"
INTEGER value "suitability 0-100"
}
metric {
INTEGER metric_seq PK "auto-increment"
VARCHAR metric_key "metric identifier"
VARCHAR description
}
cell_metric {
INTEGER cell_id FK "-> cell"
INTEGER metric_seq FK "-> metric"
DOUBLE value
}
zone {
INTEGER zone_seq PK "auto-increment"
VARCHAR tbl "source table"
VARCHAR fld "source field"
VARCHAR value "zone value"
}
zone_cell {
INTEGER zone_seq FK "-> zone"
INTEGER cell_id FK "-> cell"
INTEGER pct_covered "% cell in zone"
}
zone_metric {
INTEGER zone_seq FK "-> zone"
INTEGER metric_seq FK "-> metric"
DOUBLE value
}
taxon {
DOUBLE taxon_id PK "taxon identifier"
VARCHAR taxon_authority "worms, itis, etc."
VARCHAR scientific_name
VARCHAR common_name
VARCHAR redlist_code
VARCHAR extrisk_code
BOOLEAN is_ok
}
taxon_model {
DOUBLE taxon_id FK "-> taxon"
VARCHAR ds_key FK "-> dataset"
INTEGER mdl_seq FK "-> model"
}
zone_taxon {
VARCHAR zone_tbl
VARCHAR zone_fld
VARCHAR zone_value
DOUBLE taxon_id FK "-> taxon"
INTEGER mdl_seq FK "-> model"
DOUBLE suit_rl_area "sensitivity score"
}
listing {
DOUBLE spp_id
DOUBLE worms_id
DOUBLE botw_id
VARCHAR extrisk_code
INTEGER er_score
BOOLEAN is_mmpa
}
dataset ||--|{ model : ds_key
dataset ||--|{ species : ds_key
model ||--|{ model_cell : mdl_seq
model ||--|{ taxon_model : mdl_seq
cell ||--|{ model_cell : cell_id
cell ||--|{ cell_metric : cell_id
cell ||--|{ zone_cell : cell_id
metric ||--|{ cell_metric : metric_seq
metric ||--|{ zone_metric : metric_seq
zone ||--|{ zone_cell : zone_seq
zone ||--|{ zone_metric : zone_seq
taxon ||--|{ taxon_model : taxon_id
taxon ||--|{ zone_taxon : taxon_id
```
## Tables
```{r}
#| label: sdm-tables
#| output: asis
#| echo: false
sdm_tables <- sort(dbListTables(con_sdm))
src <- map_chr(sdm_tables, function(tbl_name) {
knitr::knit_child(
text = c(
glue("### {tbl_name}\n"),
"```{r}",
glue('#| label: sdm-tbl-{tbl_name}'),
"#| echo: false",
glue('tbl_info(con_sdm, "{tbl_name}")'),
"```",
""
),
envir = environment(),
quiet = TRUE
)
})
cat("::: {.panel-tabset}\n\n")
cat(src, sep = "\n")
cat("\n:::\n")
```
# SPP — Species Taxonomic Authority
```{r}
#| label: spp-changes
#| echo: false
#| output: asis
check_schema(con_spp, spp_expected, "SPP")
```
## Schema
```{mermaid}
%%| label: fig-spp-erd
%%| fig-cap: "SPP entity relationship diagram (click to zoom)"
%%| fig-width: 10
erDiagram
worms {
INTEGER taxonID PK "WoRMS taxon id"
VARCHAR scientificName "scientific name"
VARCHAR taxonRank "rank"
VARCHAR taxonomicStatus "accepted, etc."
VARCHAR kingdom
VARCHAR phylum
VARCHAR class
VARCHAR family
BOOLEAN isMarine
BOOLEAN isExtinct
}
worms_vernacular {
INTEGER taxonID FK "-> worms"
VARCHAR vernacularName "common name"
VARCHAR language
}
gbif {
INTEGER taxonID PK "GBIF taxon id"
VARCHAR canonicalName "canonical name"
VARCHAR taxonRank
VARCHAR taxonomicStatus
VARCHAR kingdom
VARCHAR phylum
VARCHAR class
VARCHAR family
}
gbif_vernacular {
DOUBLE taxonID FK "-> gbif"
VARCHAR vernacularName "common name"
VARCHAR language
VARCHAR country
}
itis {
INTEGER taxonID PK "ITIS taxon id"
VARCHAR scientificName
VARCHAR taxonRank
VARCHAR taxonomicStatus
}
itis_vernacular {
INTEGER taxonID FK "-> itis"
VARCHAR vernacularName "common name"
VARCHAR language
}
botw {
DOUBLE taxonID PK "BirdLife id"
VARCHAR scientificName
VARCHAR redlist_code "IUCN code"
VARCHAR kingdom
VARCHAR family
}
botw_vernacular {
DOUBLE taxonID FK "-> botw"
VARCHAR vernacularName "common name"
BOOLEAN isPreferredName
}
iucn_redlist {
INTEGER sis_taxon_id PK "IUCN SIS taxon id"
VARCHAR taxon_scientific_name "scientific name"
VARCHAR red_list_category_code "IUCN code"
INTEGER assessment_id
BOOLEAN latest
}
iucn_vernacular {
DOUBLE taxonID FK "-> iucn_redlist"
VARCHAR vernacularName "common name"
VARCHAR language
BOOLEAN isPreferredName
}
worms ||--|{ worms_vernacular : taxonID
gbif ||--|{ gbif_vernacular : taxonID
itis ||--|{ itis_vernacular : taxonID
botw ||--|{ botw_vernacular : taxonID
iucn_redlist ||--|{ iucn_vernacular : "sis_taxon_id = taxonID"
```
## Tables
```{r}
#| label: spp-tables
#| output: asis
#| echo: false
spp_tables <- sort(dbListTables(con_spp))
src <- map_chr(spp_tables, function(tbl_name) {
knitr::knit_child(
text = c(
glue("### {tbl_name}\n"),
"```{r}",
glue('#| label: spp-tbl-{tbl_name}'),
"#| echo: false",
glue('tbl_info(con_spp, "{tbl_name}")'),
"```",
""
),
envir = environment(),
quiet = TRUE
)
})
cat("::: {.panel-tabset}\n\n")
cat(src, sep = "\n")
cat("\n:::\n")
```
```{r}
#| label: cleanup
#| include: false
dbDisconnect(con_sdm, shutdown = TRUE)
dbDisconnect(con_spp, shutdown = TRUE)
```