---
title: "GoA Marine Mammal Species Review"
subtitle: "Response to SEFSC feedback on 12 species in Eastern Gulf (v2 → v6)"
format:
html:
code-fold: true
code-tools: true
editor_options:
chunk_output_type: console
---
## Executive Summary
NOAA SEFSC flagged 12 marine mammal species that the Marine Sensitivity Tool
(MST) showed as occurring in the Eastern Gulf of America but shouldn't be
there. **As of v6, all 12 species are resolved** — none remain erroneously
in the Gulf:
| Resolution | # | Species | Mechanism |
|---|---|---|---|
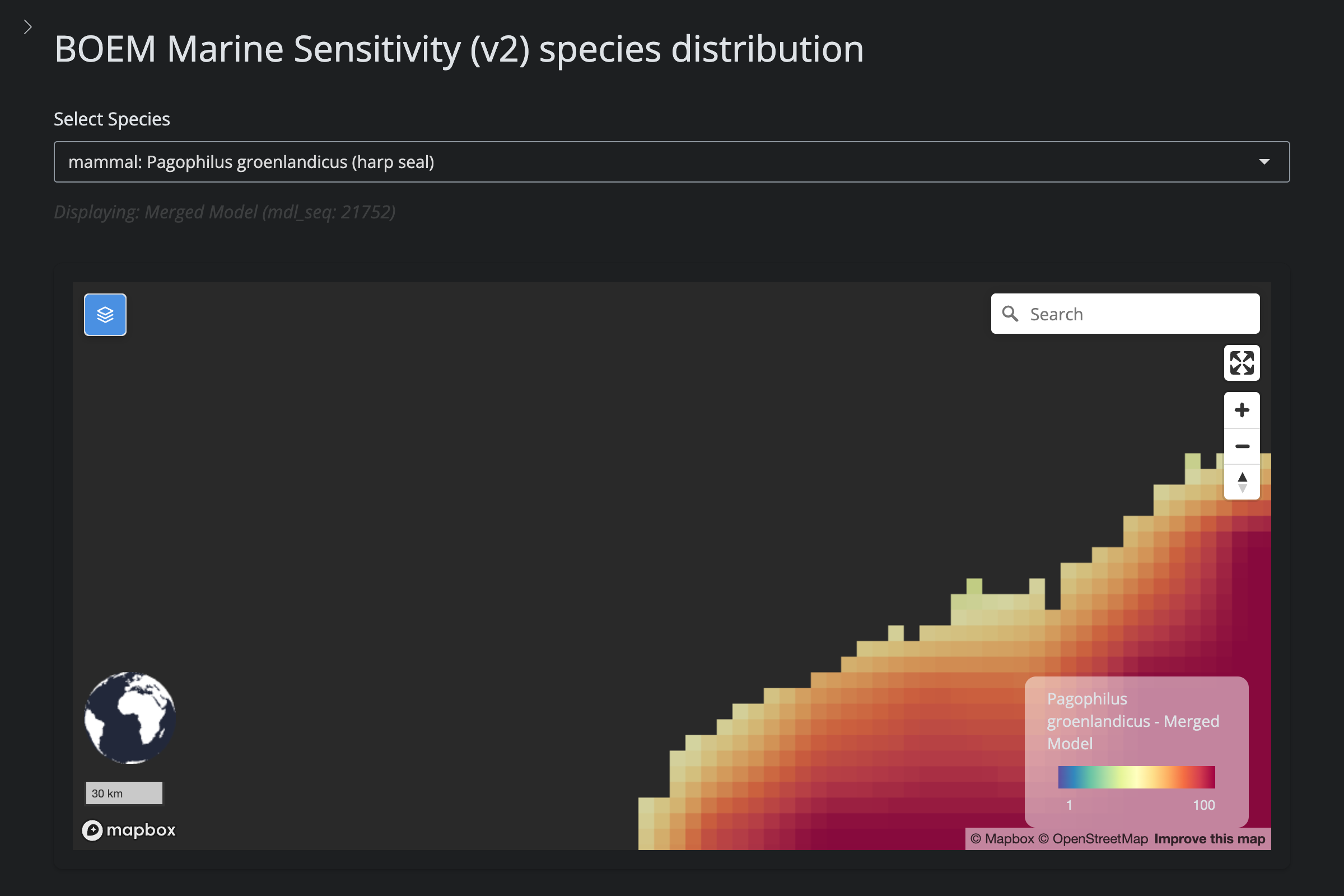
| **Removed from GOA (v5)** | 7 | Hooded Seal, Long-Finned Pilot Whale, Gray Seal, Northern Bottlenose Whale, White-Beaked Dolphin, Sowerby's Beaked Whale, Harp Seal | IUCN range mask clipped AquaMaps to expert range (outside Gulf) |
| **Excluded as duplicate (v5)** | 1 | *B. brydei* | Taxonomic synonym of *B. edeni*; `is_ok=FALSE` |
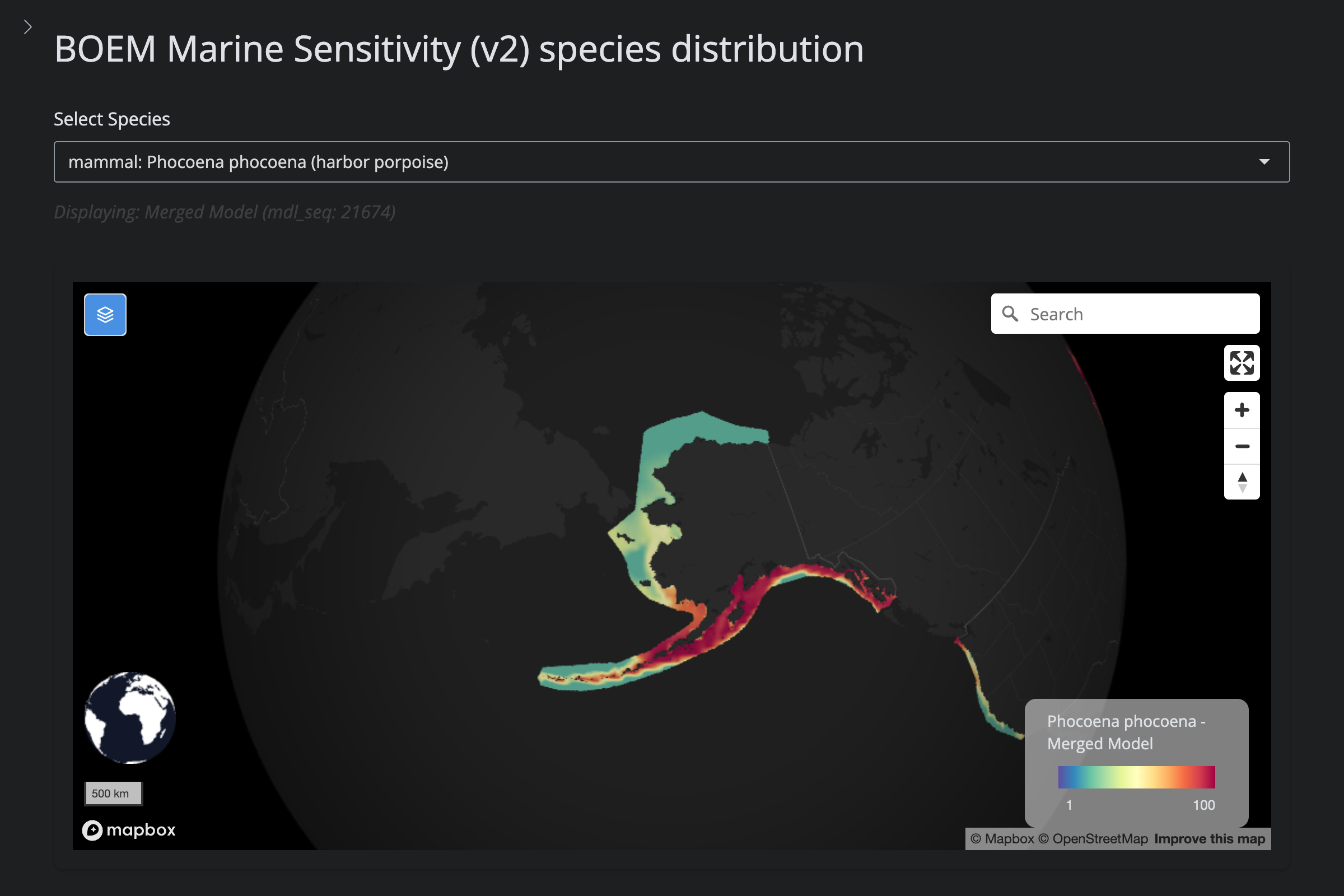
| **Reduced GOA footprint (v5)** | 2 | *B. edeni* (GAA+GAB only), *P. phocoena* (GOA only) | IUCN range mask trimmed to actual Gulf extent |
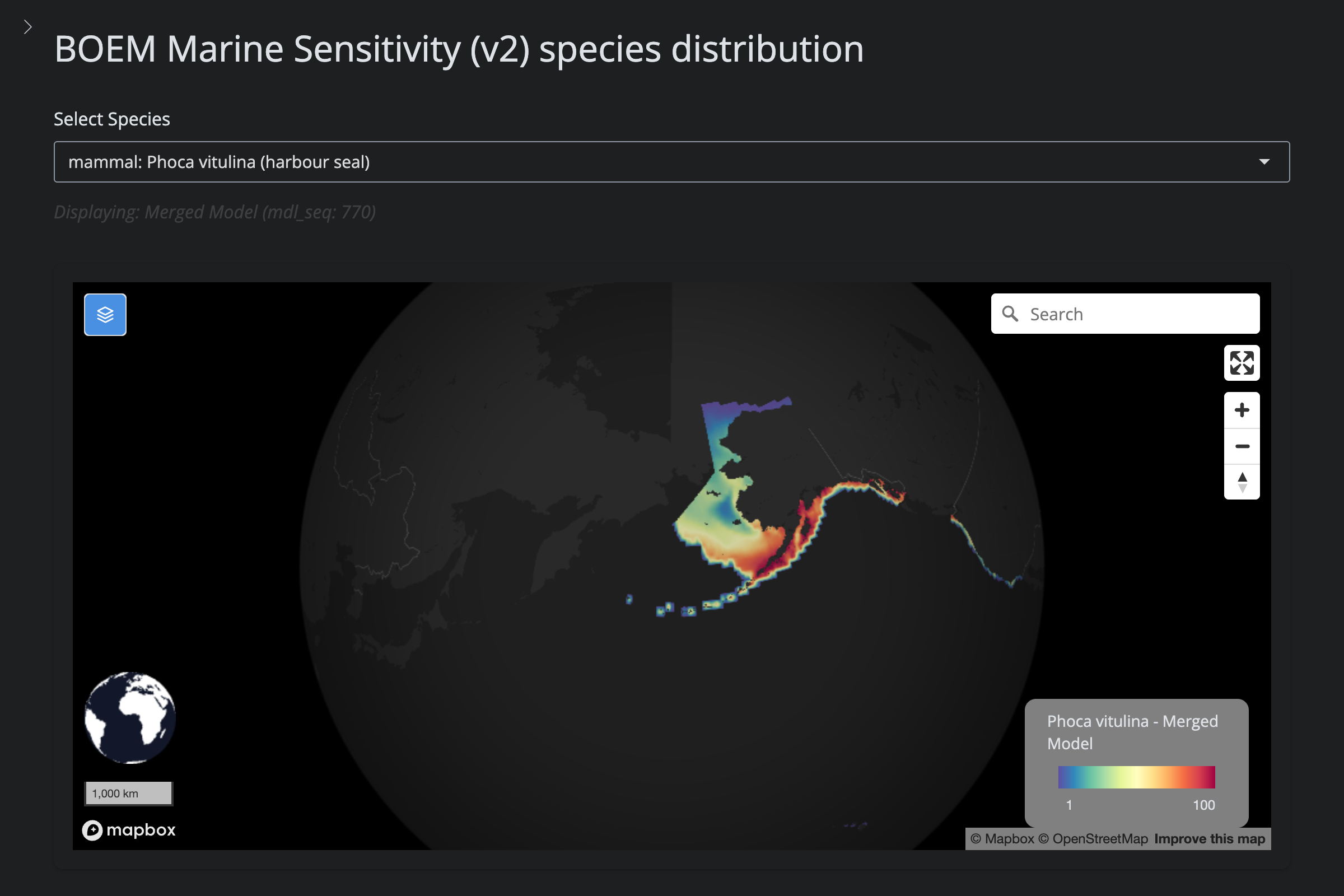
| **Removed from GOA (v6)** | 1 | Harbor Seal (*Phoca vitulina*) | IUCN range from newer `MAMMALS.zip` download; range is Atlantic/Pacific coasts only |
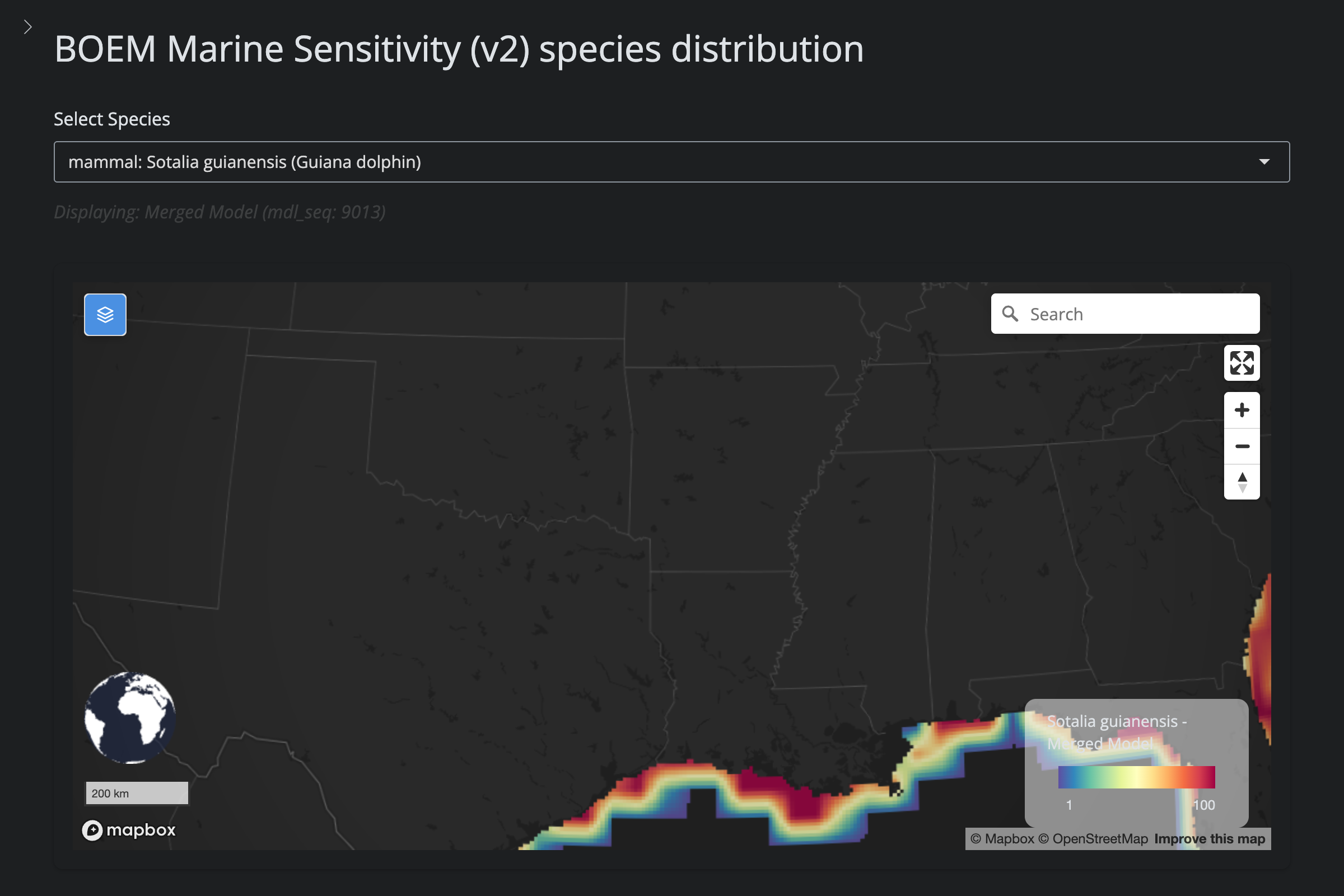
| **Excluded — range outside EEZ (v6)** | 1 | Guiana Dolphin (*Sotalia guianensis*) | IUCN range produces 0 cells in US EEZ; merged model truncated, `is_ok=FALSE` |
The two species that remained problematic through v5 were resolved in v6 by:
1. Ingesting 3 missing mammal IUCN ranges from a newer `MAMMALS.zip` download
(the original `MAMMALS_MARINE_ONLY.zip` and
`MAMMALS_MARINE_AND_TERRESTRIAL.zip` were incomplete).
2. Excluding 371 species (343 fish, 20 invertebrates, 6 corals, 2 mammals)
whose IUCN ranges fall entirely outside the US EEZ — their AquaMaps
predictions in US waters were edge-of-range artifacts with near-zero
suitability.
See [investigate_iucn_masking.qmd](investigate_iucn_masking.qmd) for the
broader impact analysis across all taxa groups.
```{r}
#| label: setup
#| warning: false
librarian::shelf(
DBI,
dplyr,
duckdb,
DT,
fs,
glue,
here,
htmltools,
knitr,
purrr,
readr,
stringr,
tibble,
tidyr,
webshot2,
quiet = T)
knitr::opts_chunk$set(echo = TRUE, warning = FALSE)
source(here("libs/paths.R"))
# v6 database (current version, from paths.R)
con_sdm <- dbConnect(duckdb(dbdir = sdm_db, read_only = TRUE))
# v2 database (outside dir_big/derived on laptop)
sdm_db_v2 <- ifelse(
is_server,
"/share/data/big/v2/sdm.duckdb",
"~/_big/msens/v2/sdm.duckdb")
con_sdm_v2 <- dbConnect(duckdb(dbdir = sdm_db_v2, read_only = TRUE))
# output directory for screenshots
dir_figs <- here("figs/explore_goa-marmam")
dir_create(dir_figs)
# goa program area codes
goa_zones <- c("GAA", "GAB", "GOA")
# zone table names per version
tbl_pra_v2 <- "ply_programareas_2026"
# tbl_pra (v6) already defined by paths.R
```
## Background
NOAA SEFSC (Avery Paxton, 2026-01-29) flagged 12 marine mammal species:
> One of my action items was to share the list of other marine mammals that the
> MST shows as occurring in the Eastern Gulf planning area (that's where we
> focused our initial review) that don't currently occur there. There were 12
> more (13 if include walrus) of 44 marine mammals.
**References:**
- [GitHub apps#5 comment](https://github.com/MarineSensitivity/apps/issues/5#issuecomment-4118545339)
— Harbor seal IUCN range map issue
- [Google Doc notes](https://docs.google.com/document/d/1s6HWNkyethIL5-VjE3tBGu9o_YIu3oFcEutw7ouN1Bs/edit?usp=sharing)
- [SEFSC meeting notes](dev/2026-02-05_v3.md)
- [investigate_iucn_masking.qmd](investigate_iucn_masking.qmd)
— quantifies impact of 0-cell IUCN masking across all taxa groups
## Resolution Table
```{r}
#| label: resolution_table
d_spp <- tibble::tribble(
~scientific_name, ~common_name, ~v2_mdl_seq,
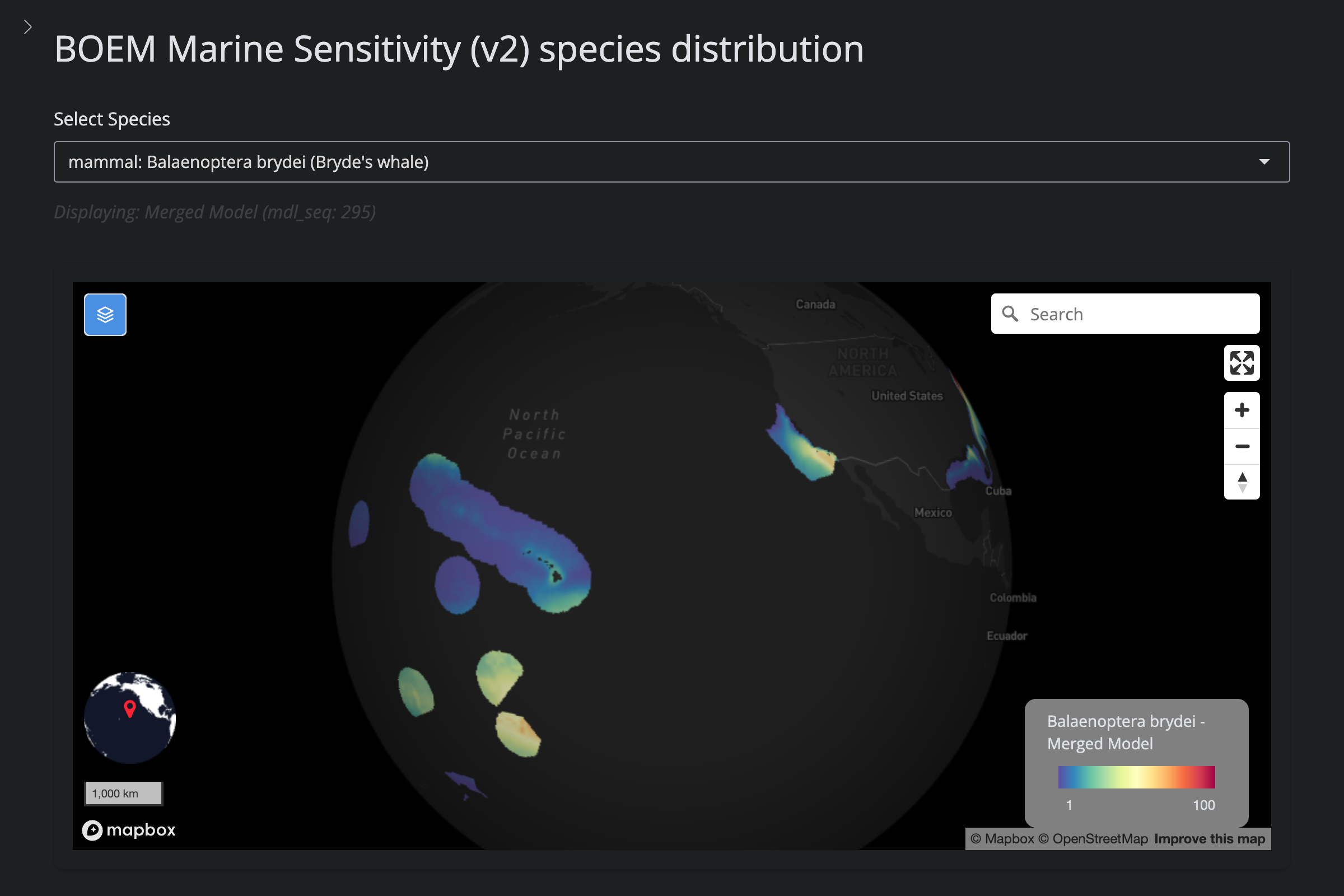
"Balaenoptera brydei", "Bryde's whale", 295L,
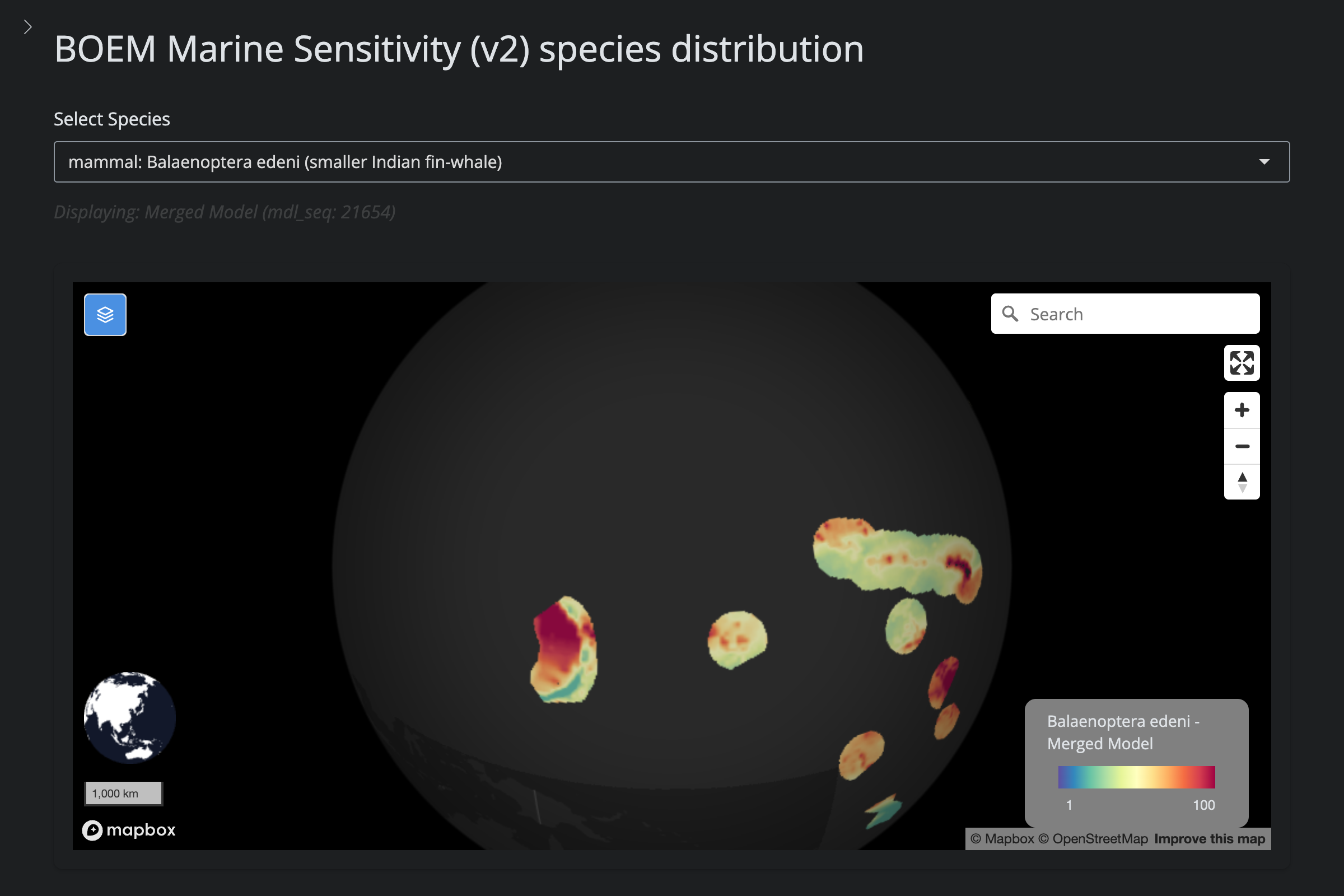
"Balaenoptera edeni", "Bryde's Whale", 170L,
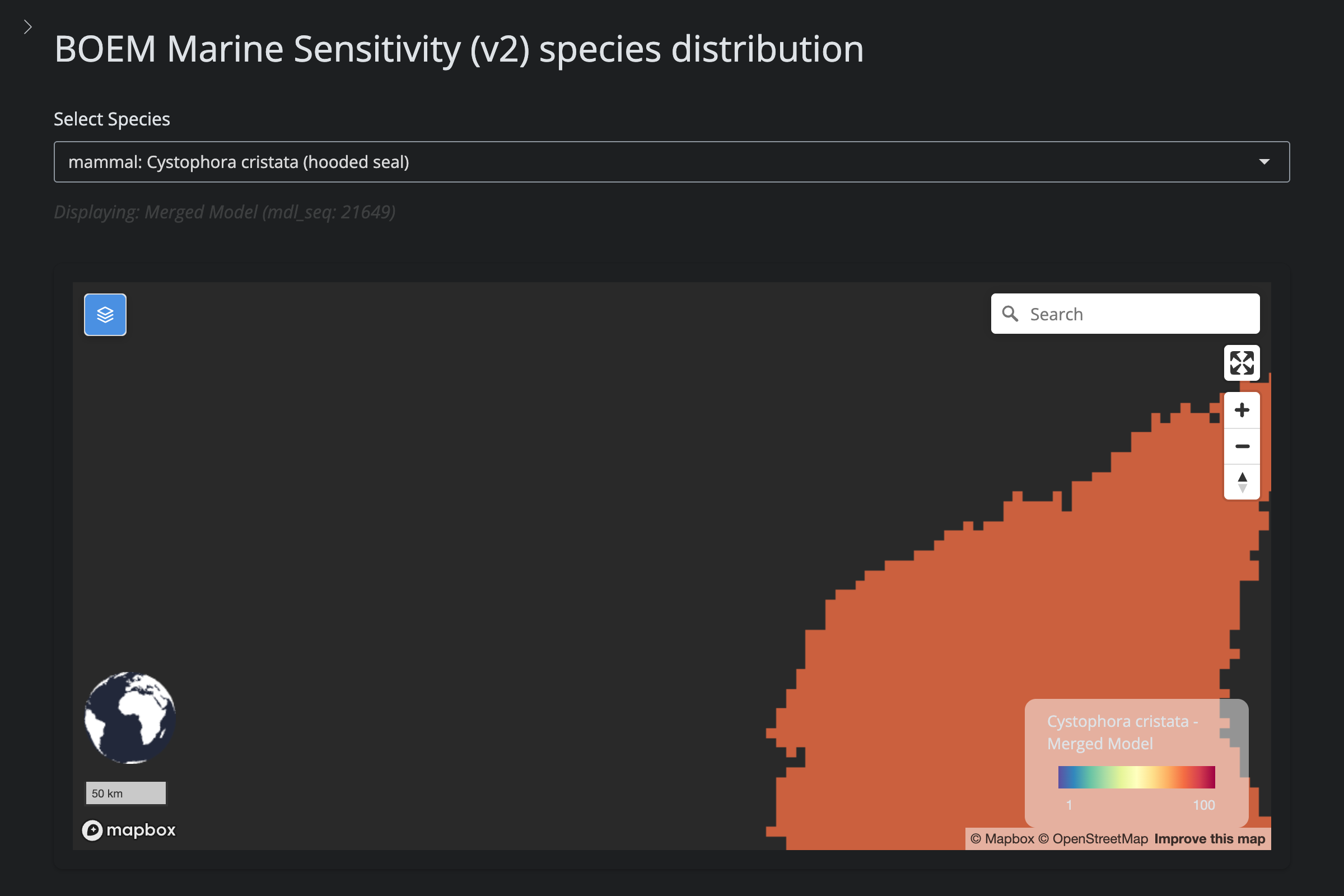
"Cystophora cristata", "Hooded Seal", 1808L,
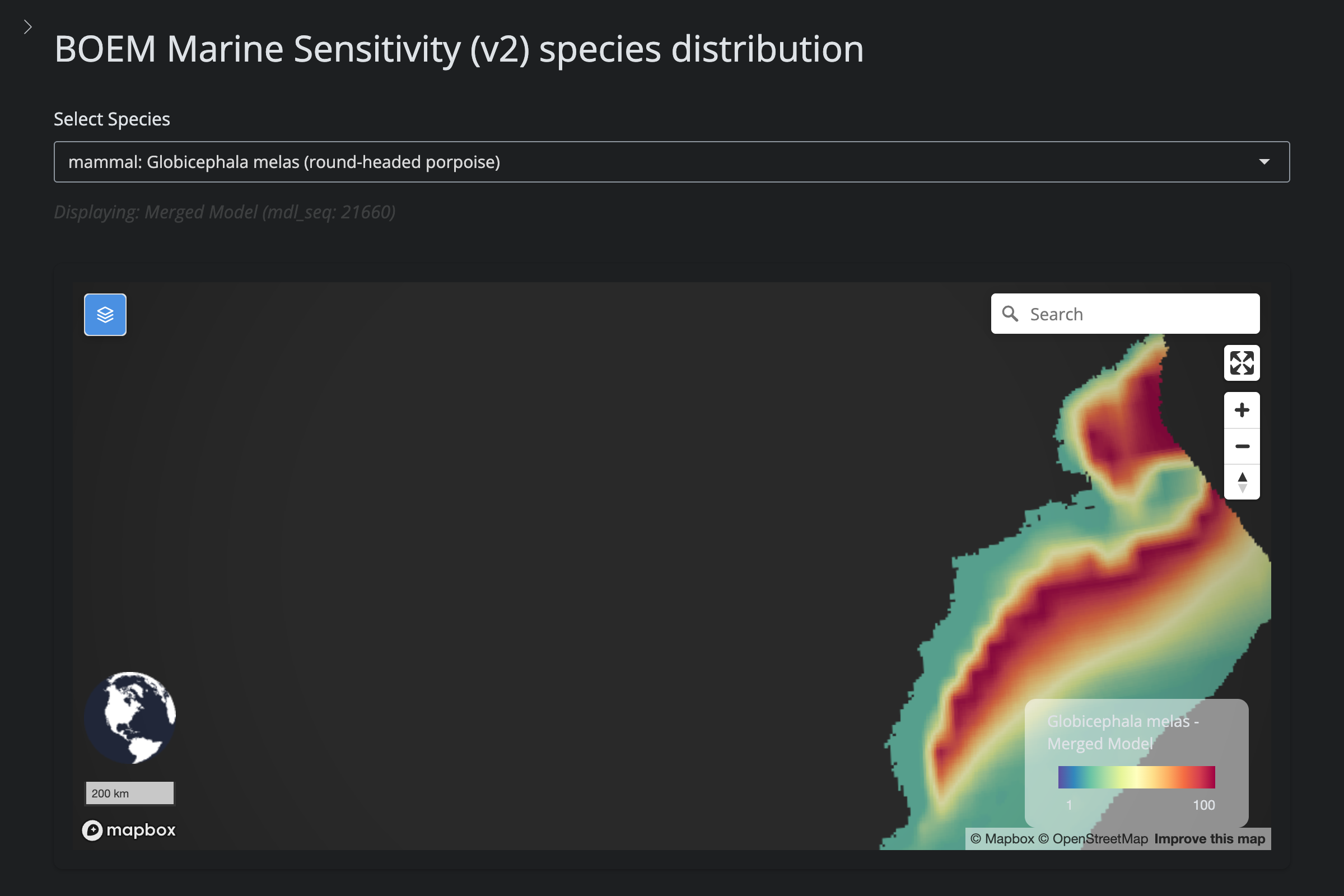
"Globicephala melas", "Long-Finned Pilot Whale", 1434L,
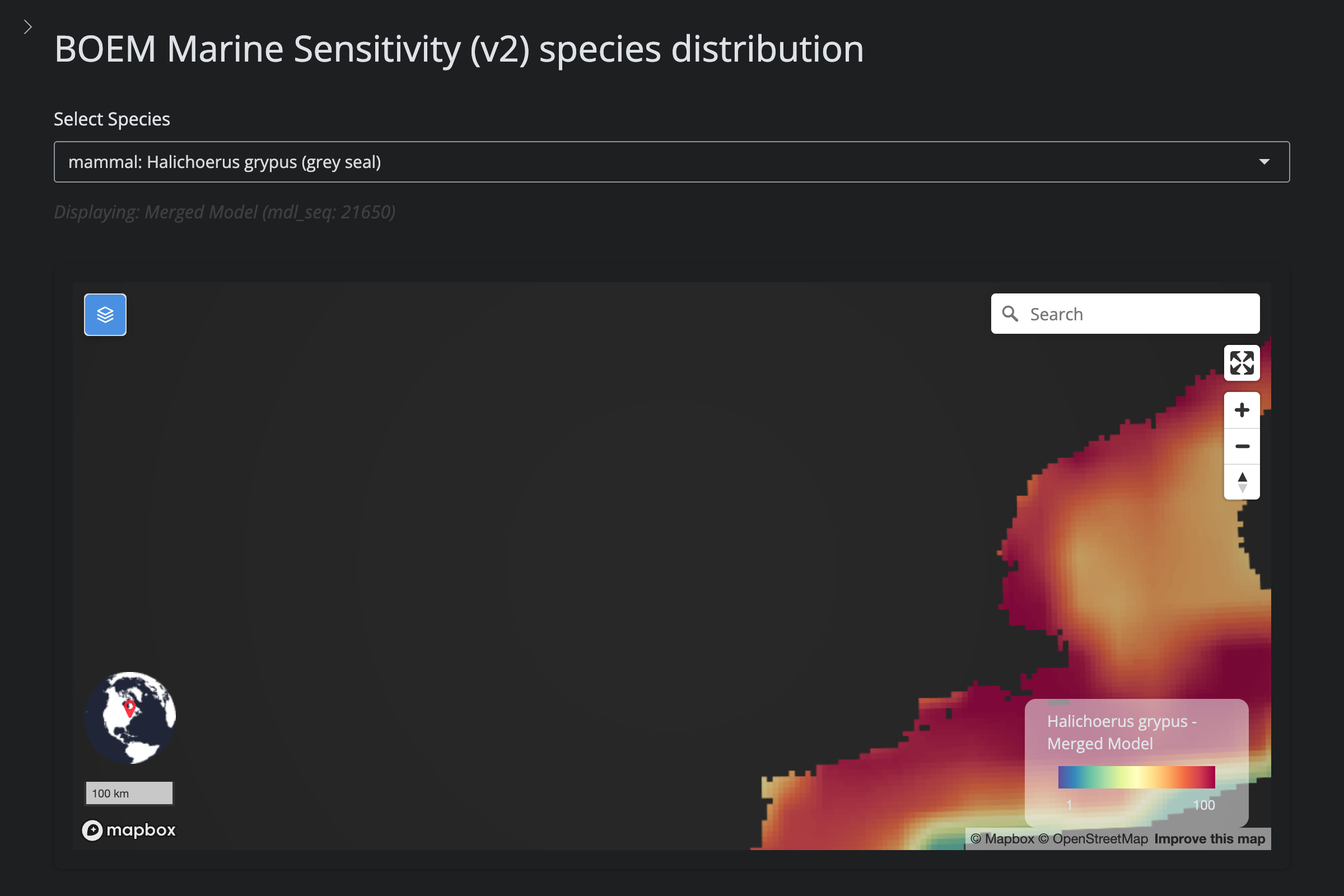
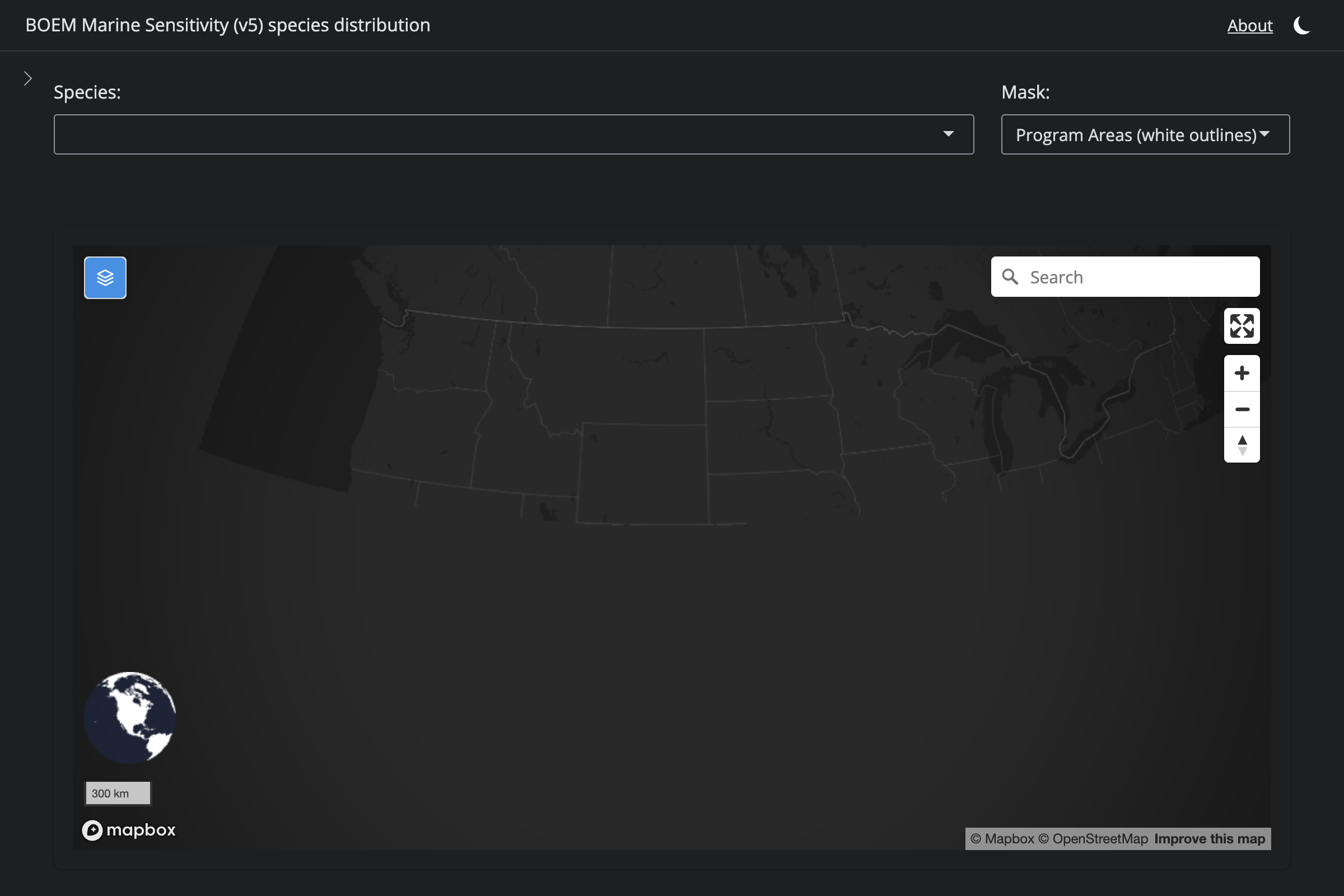
"Halichoerus grypus", "Gray Seal", 4454L,
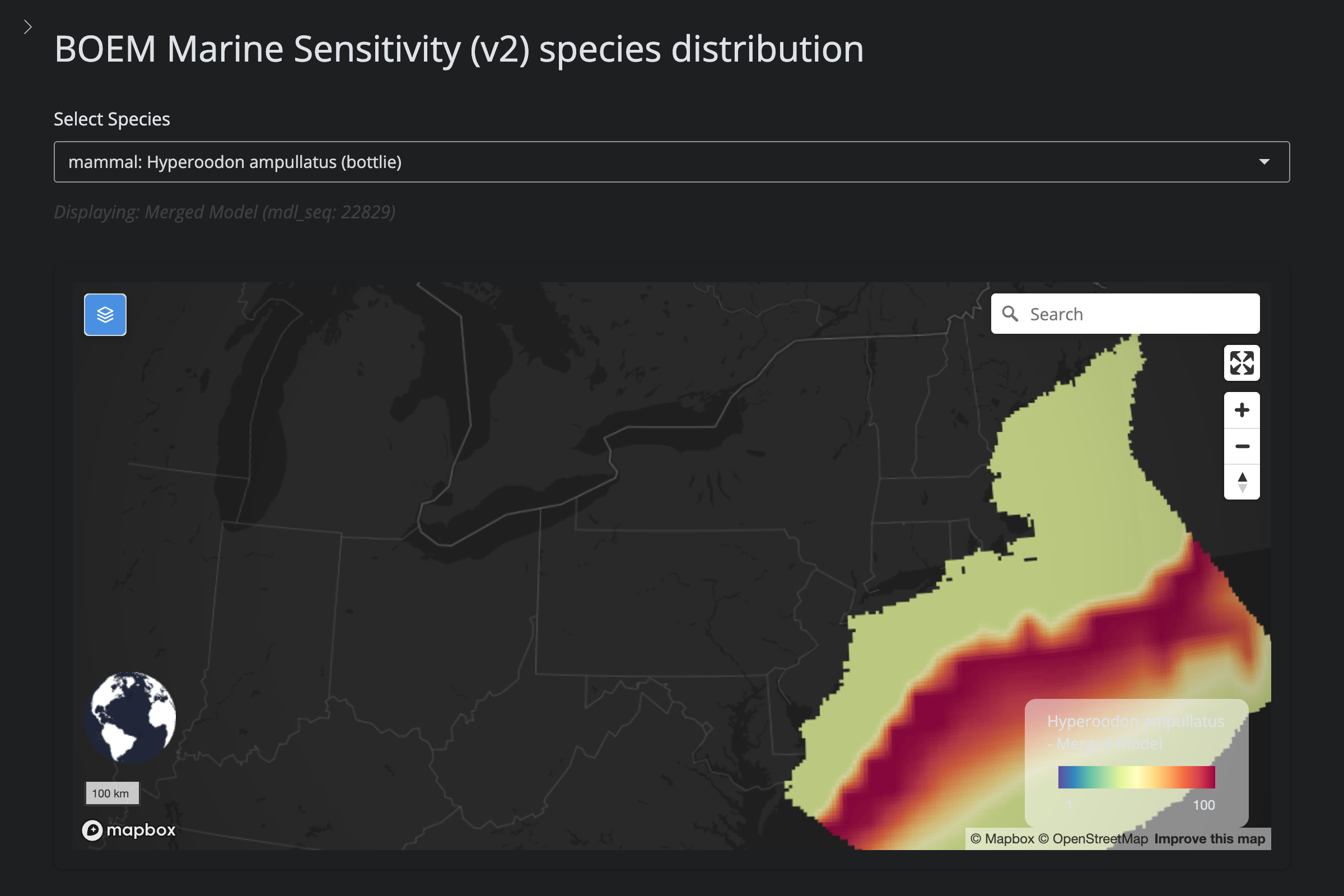
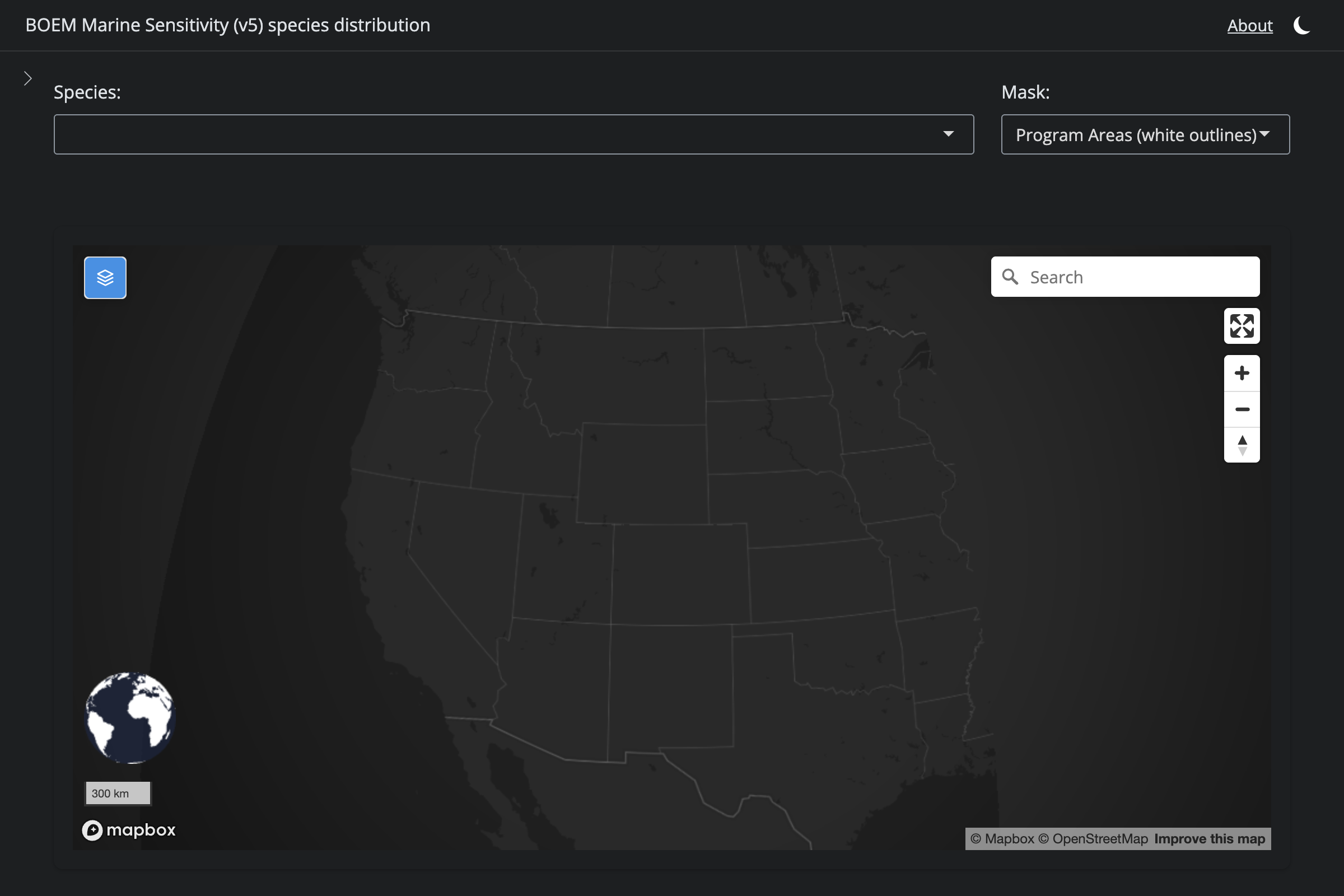
"Hyperoodon ampullatus", "Northern Bottlenose Whale", 1354L,
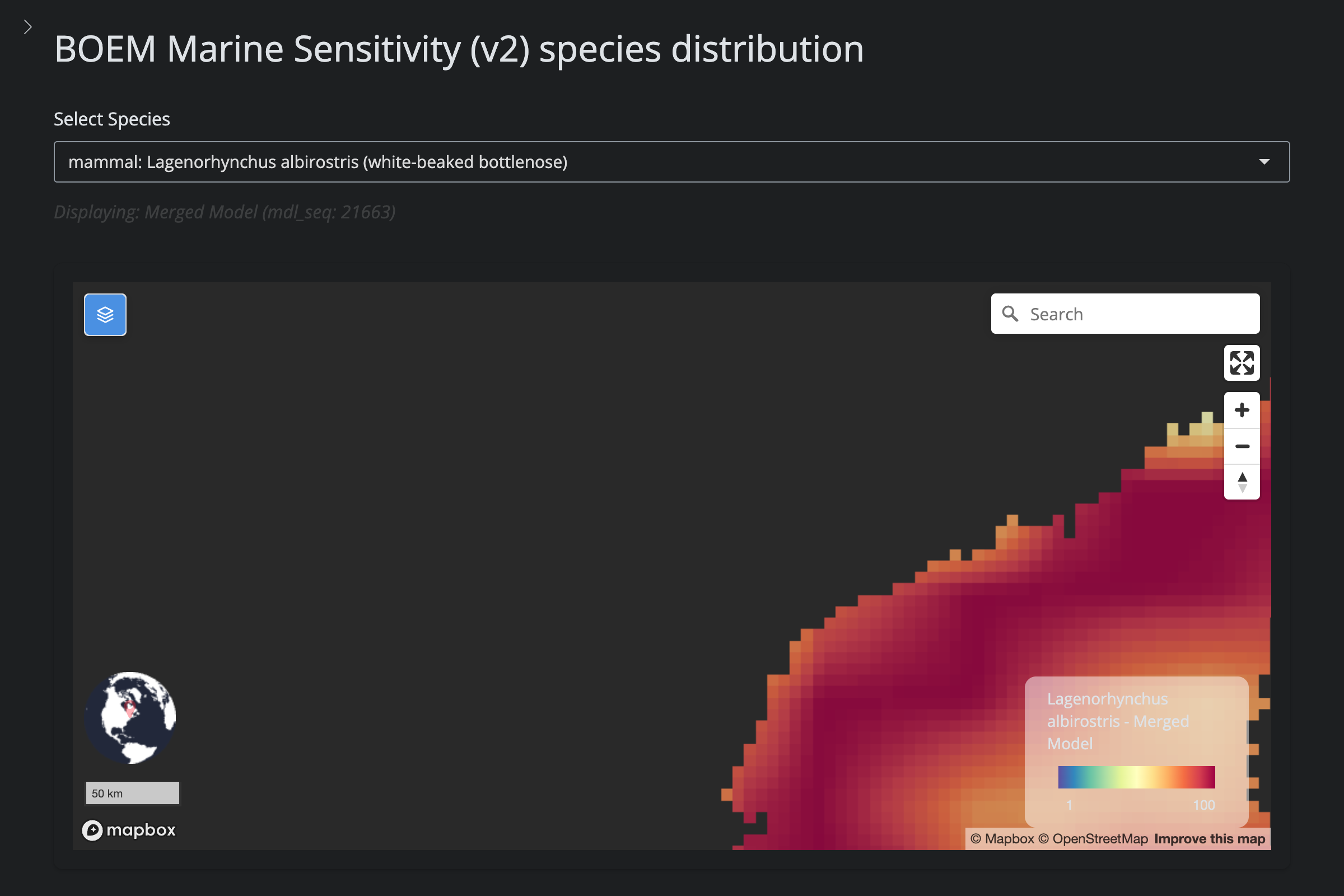
"Lagenorhynchus albirostris", "White-Beaked Dolphin", 1679L,
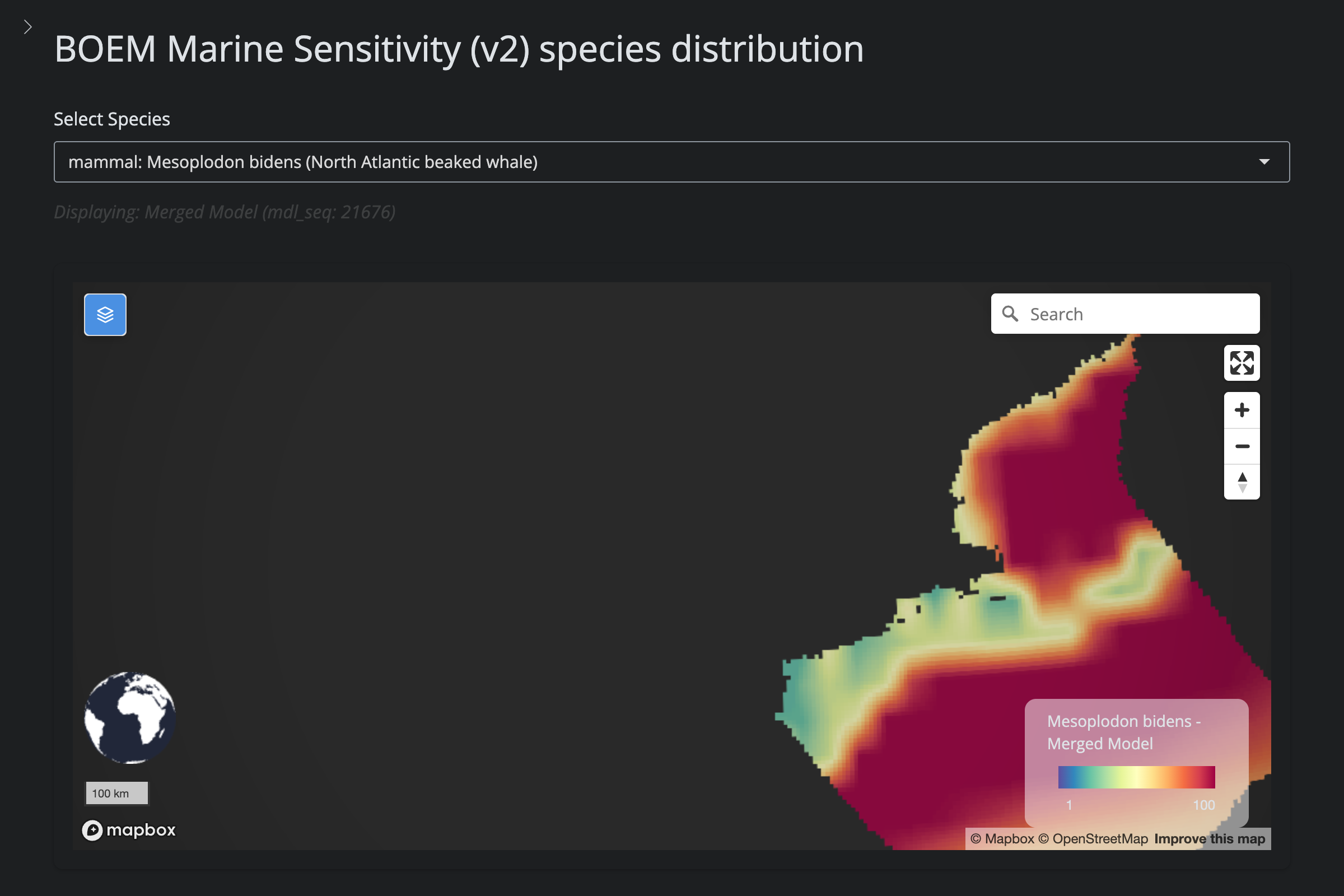
"Mesoplodon bidens", "Sowerby's Beaked Whale", 1308L,
"Pagophilus groenlandicus", "Harp Seal", 3246L,
"Phoca vitulina", "Harbor Seal", 770L,
"Phocoena phocoena", "Harbor Porpoise", 593L,
"Sotalia guianensis", "Guiana dolphin", 9013L)
# look up v6 taxon info
d_taxon_spp <- tbl(con_sdm, "taxon") |>
filter(scientific_name %in% !!d_spp$scientific_name) |>
collect()
d_spp <- d_spp |>
left_join(
d_taxon_spp |>
select(
scientific_name, taxon_id,
v6_mdl_seq = mdl_seq,
is_ok, is_mmpa, er_score,
extrisk_code, redlist_code, n_ds),
by = "scientific_name")
# v2 and v6 GOA zone_taxon
d_goa_v2 <- dbGetQuery(con_sdm_v2, glue(
"SELECT sp_scientific, zone_value, avg_suit, rl_score, suit_rl, area_km2
FROM zone_taxon
WHERE zone_tbl = '{tbl_pra_v2}'
AND zone_value IN ('GAA', 'GAB', 'GOA')
AND sp_scientific IN ({paste0(\"'\", d_spp$scientific_name, \"'\", collapse = ', ')})
ORDER BY sp_scientific, zone_value")) |>
as_tibble()
d_goa_v6 <- dbGetQuery(con_sdm, glue(
"SELECT sp_scientific, zone_value, avg_suit, er_score, suit_rl, area_km2
FROM zone_taxon
WHERE zone_tbl = '{tbl_pra}'
AND zone_value IN ('GAA', 'GAB', 'GOA')
AND sp_scientific IN ({paste0(\"'\", d_spp$scientific_name, \"'\", collapse = ', ')})
ORDER BY sp_scientific, zone_value")) |>
as_tibble()
# rng_iucn cell counts per species
d_iucn_cells <- tbl(con_sdm, "taxon_model") |>
filter(
ds_key == "rng_iucn",
taxon_id %in% !!d_spp$taxon_id) |>
left_join(
tbl(con_sdm, "model_cell") |>
group_by(mdl_seq) |>
summarize(n_iucn_cells = n(), .groups = "drop"),
by = "mdl_seq") |>
collect() |>
mutate(n_iucn_cells = coalesce(n_iucn_cells, 0L)) |>
select(taxon_id, n_iucn_cells)
# build resolution table
spp_in_goa_v6 <- d_goa_v6 |> distinct(sp_scientific) |> pull()
goa_zones_v2 <- d_goa_v2 |>
group_by(sp_scientific) |>
summarize(goa_zones_v2 = paste(sort(zone_value), collapse = ", "), .groups = "drop")
goa_zones_v6 <- d_goa_v6 |>
group_by(sp_scientific) |>
summarize(goa_zones_v6 = paste(sort(zone_value), collapse = ", "), .groups = "drop")
d_resolution <- d_spp |>
left_join(d_iucn_cells, by = "taxon_id") |>
left_join(goa_zones_v2, by = c("scientific_name" = "sp_scientific")) |>
left_join(goa_zones_v6, by = c("scientific_name" = "sp_scientific")) |>
mutate(
in_goa_v2 = scientific_name %in% d_goa_v2$sp_scientific,
in_goa_v6 = scientific_name %in% spp_in_goa_v6,
resolution = case_when(
scientific_name == "Balaenoptera brydei" ~
"excluded (v5): taxonomic duplicate of B. edeni",
scientific_name == "Sotalia guianensis" ~
"excluded (v6): IUCN range outside US EEZ (0 cells)",
scientific_name == "Phoca vitulina" ~
"removed from GOA (v6): IUCN range ingested from MAMMALS.zip; Atlantic/Pacific only",
scientific_name == "Balaenoptera edeni" ~
"reduced (v5): IUCN range limits to GAA+GAB (no GOA)",
scientific_name == "Phocoena phocoena" ~
"reduced (v5): IUCN range limits to GOA only",
!is_ok & !in_goa_v6 ~
"removed from GOA (v5): IUCN range mask outside Gulf",
in_goa_v2 & !in_goa_v6 ~
"removed from GOA (v5): IUCN range mask outside Gulf",
.default = "unchanged")) |>
select(
scientific_name, common_name,
is_ok, n_iucn_cells,
goa_zones_v2, goa_zones_v6,
resolution) |>
arrange(resolution, scientific_name)
datatable(
d_resolution,
caption = "resolution status for all 12 SEFSC-flagged species in v6",
escape = FALSE,
filter = "top",
options = list(dom = "ft", pageLength = 15, scrollX = TRUE))
```
## Database Interrogation (v6)
```{r}
#| label: db_interrogate
# all models per species (per-dataset and merged)
d_models_spp <- tbl(con_sdm, "taxon_model") |>
filter(taxon_id %in% !!d_taxon_spp$taxon_id) |>
collect() |>
left_join(
d_taxon_spp |> select(taxon_id, scientific_name, common_name),
by = "taxon_id")
# cell counts per model (left_join: 0-cell rng_iucn models have no model_cell rows)
d_counts_spp <- tbl(con_sdm, "model_cell") |>
filter(mdl_seq %in% !!d_models_spp$mdl_seq) |>
group_by(mdl_seq) |>
summarize(
n_cells = n(),
v_min = min(value, na.rm = TRUE),
v_max = max(value, na.rm = TRUE),
.groups = "drop") |>
collect()
d_summary_spp <- d_models_spp |>
left_join(d_counts_spp, by = "mdl_seq") |>
mutate(n_cells = coalesce(n_cells, 0L)) |>
select(scientific_name, common_name, ds_key, mdl_seq, n_cells, v_min, v_max) |>
arrange(scientific_name, ds_key)
datatable(
d_summary_spp,
caption = "v6 models per species by dataset (n_cells=0 means IUCN range outside US EEZ)",
filter = "top",
options = list(dom = "ft", pageLength = 50))
```
### Dataset summary (wide)
```{r}
#| label: ds_wide
d_ds_wide <- d_summary_spp |>
select(scientific_name, ds_key, n_cells) |>
pivot_wider(
names_from = ds_key,
values_from = n_cells,
names_prefix = "n_")
d_taxon_wide <- d_taxon_spp |>
select(
scientific_name, common_name, sp_cat,
er_score, is_mmpa, is_ok,
extrisk_code, redlist_code, n_ds) |>
left_join(d_ds_wide, by = "scientific_name")
datatable(
d_taxon_wide,
caption = "v6 species summary with cell counts per dataset",
filter = "top",
options = list(dom = "ft", scrollX = TRUE))
```
## GOA Scores: v2 vs v6
```{r}
#| label: goa_comparison
d_goa_v2_r <- d_goa_v2 |>
rename(er_score_v2 = rl_score, avg_suit_v2 = avg_suit,
suit_rl_v2 = suit_rl, area_km2_v2 = area_km2)
d_goa_v6_r <- d_goa_v6 |>
rename(er_score_v6 = er_score, avg_suit_v6 = avg_suit,
suit_rl_v6 = suit_rl, area_km2_v6 = area_km2)
d_goa_cmp <- d_goa_v2_r |>
full_join(d_goa_v6_r, by = c("sp_scientific", "zone_value")) |>
left_join(
d_spp |> select(scientific_name, common_name),
by = c("sp_scientific" = "scientific_name")) |>
relocate(common_name, .after = sp_scientific) |>
arrange(sp_scientific, zone_value)
datatable(
d_goa_cmp,
caption = "GOA Program Area scores: v2 vs v6 (NA = absent from zone in that version)",
filter = "top",
options = list(dom = "ft", scrollX = TRUE, pageLength = 30))
```
## Screenshots: v2 vs v6
```{r}
#| label: webshots
for (i in 1:nrow(d_spp)) {
sp <- d_spp[i, ]
sp_slug <- str_replace_all(sp$scientific_name, " ", "-")
# v2 screenshot
url_v2 <- glue("https://app.marinesensitivity.org/mapsp_v2/?mdl_seq={sp$v2_mdl_seq}")
png_v2 <- glue("{dir_figs}/{sp_slug}_v2.png")
if (!file.exists(png_v2)) {
tryCatch(
webshot(url_v2, png_v2, vwidth = 1200, vheight = 800, zoom = 2, delay = 15),
error = function(e) message(glue("webshot failed for {sp$scientific_name} v2: {e$message}")))
}
# v6 screenshot (skip if no v6 mdl_seq)
if (!is.na(sp$v6_mdl_seq)) {
url_v6 <- glue("https://app.marinesensitivity.org/mapsp/?mdl_seq={sp$v6_mdl_seq}&splash=false")
png_v6 <- glue("{dir_figs}/{sp_slug}_v6.png")
if (!file.exists(png_v6)) {
tryCatch(
webshot(url_v6, png_v6, vwidth = 1200, vheight = 800, zoom = 2, delay = 15),
error = function(e) message(glue("webshot failed for {sp$scientific_name} v6: {e$message}")))
}
}
}
```
## Per-Species Details
```{r}
#| label: species_details
#| results: asis
#| echo: false
for (i in 1:nrow(d_spp)) {
sp <- d_spp[i, ]
sp_slug <- str_replace_all(sp$scientific_name, " ", "-")
res <- d_resolution |> filter(scientific_name == sp$scientific_name) |> pull(resolution)
cat(glue("\n\n### {sp$common_name} (*{sp$scientific_name}*) {{#{sp_slug}}}\n\n"))
if (!is.na(sp$is_ok) && !sp$is_ok) {
cat(glue(
"::: {{.callout-warning}}\n",
"**Excluded in v6** (`is_ok=FALSE`): {res}\n",
":::\n\n"))
} else {
cat(glue(
"::: {{.callout-note}}\n",
"**Resolution**: {res}\n",
":::\n\n"))
}
cat("::: {.panel-tabset}\n\n")
# tab: v2 vs v6 screenshots ----
cat("#### v2 vs v6\n\n")
cat("::: {.columns}\n")
cat("::: {.column width='50%'}\n")
url_v2 <- glue("https://app.marinesensitivity.org/mapsp_v2/?mdl_seq={sp$v2_mdl_seq}")
cat(glue("**v2** ([app link]({url_v2}))\n\n"))
png_v2 <- glue("figs/explore_goa-marmam/{sp_slug}_v2.png")
if (file.exists(here(png_v2))) {
cat(glue("\n\n"))
} else {
cat("*Screenshot not yet captured.*\n\n")
}
cat(":::\n")
cat("::: {.column width='50%'}\n")
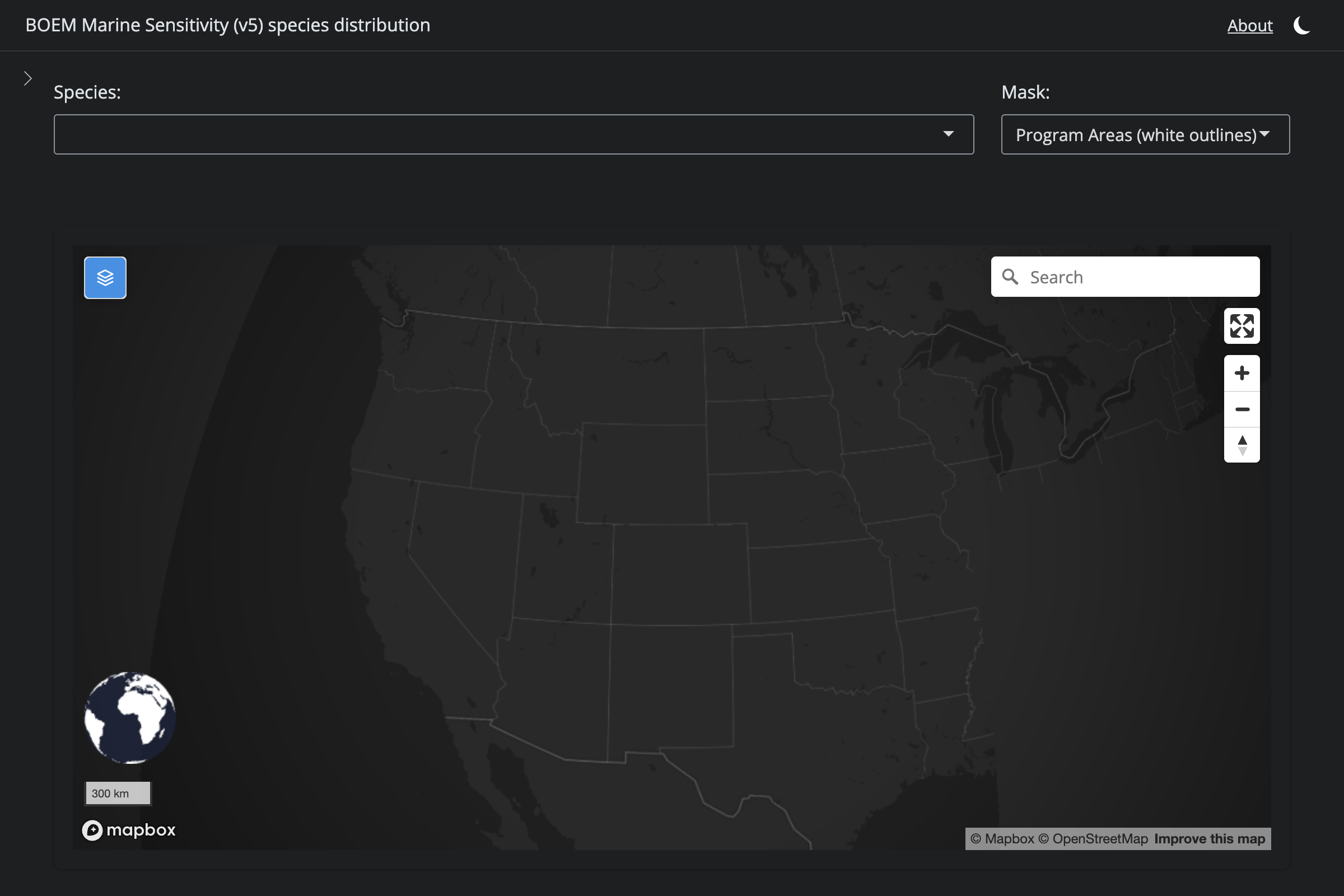
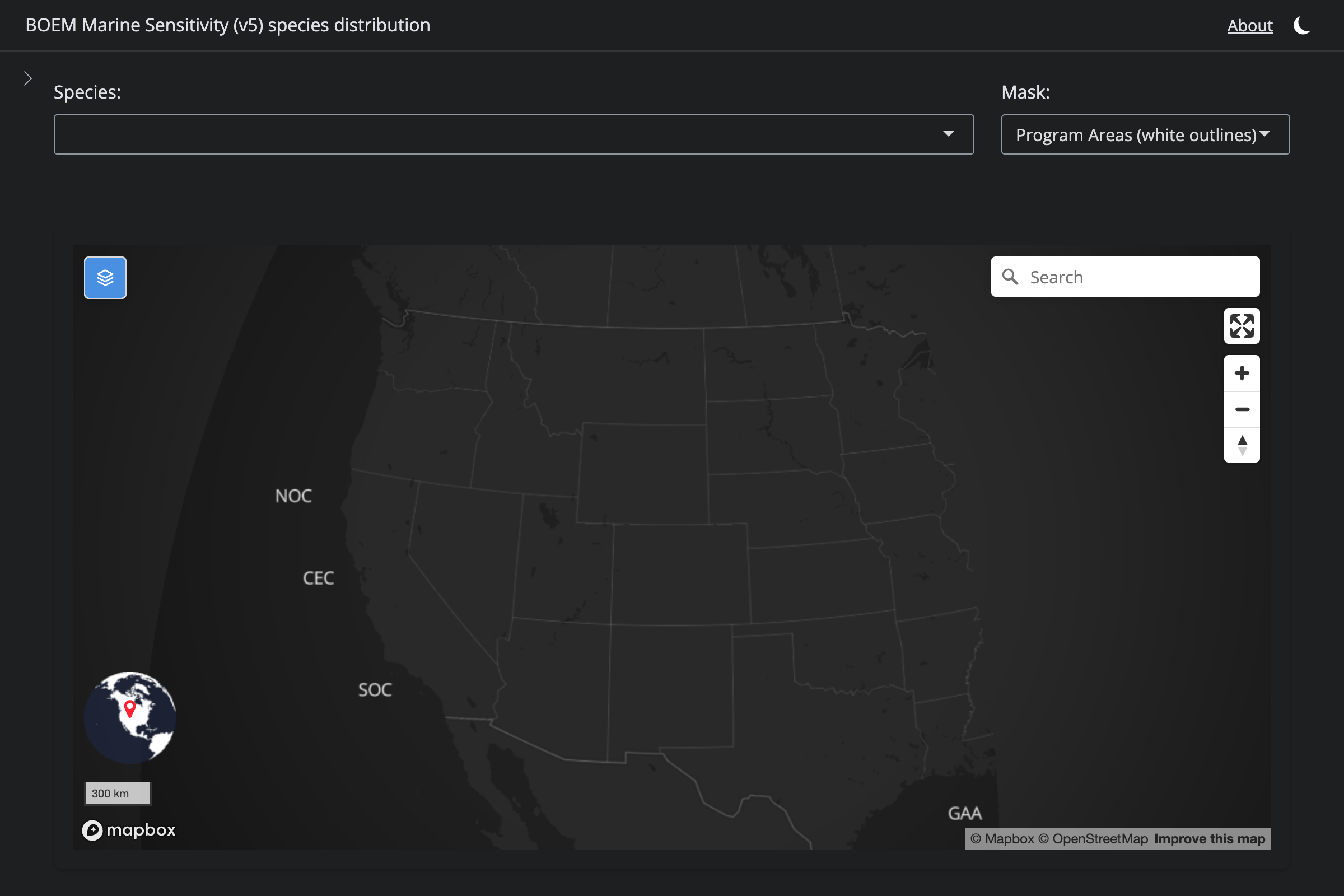
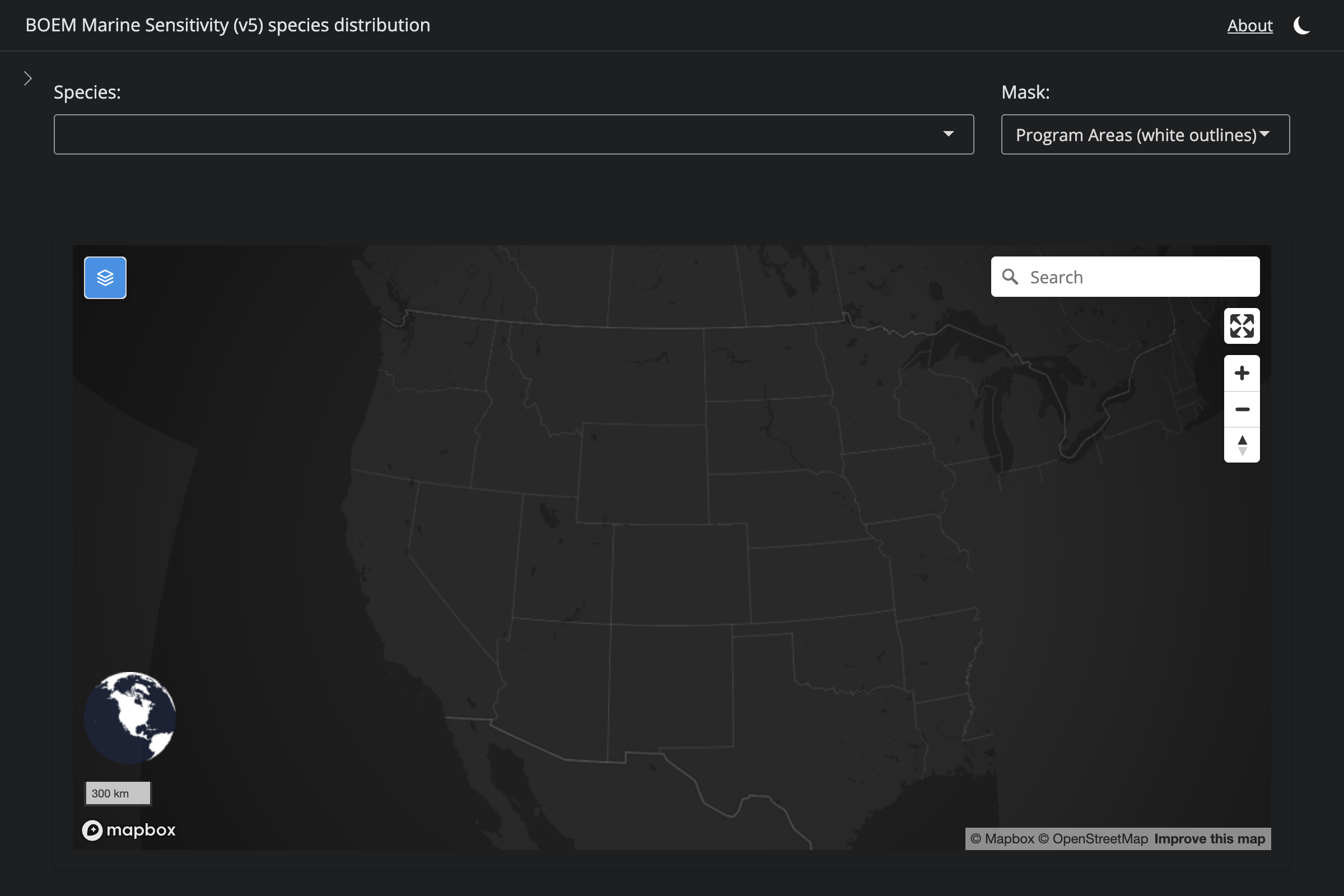
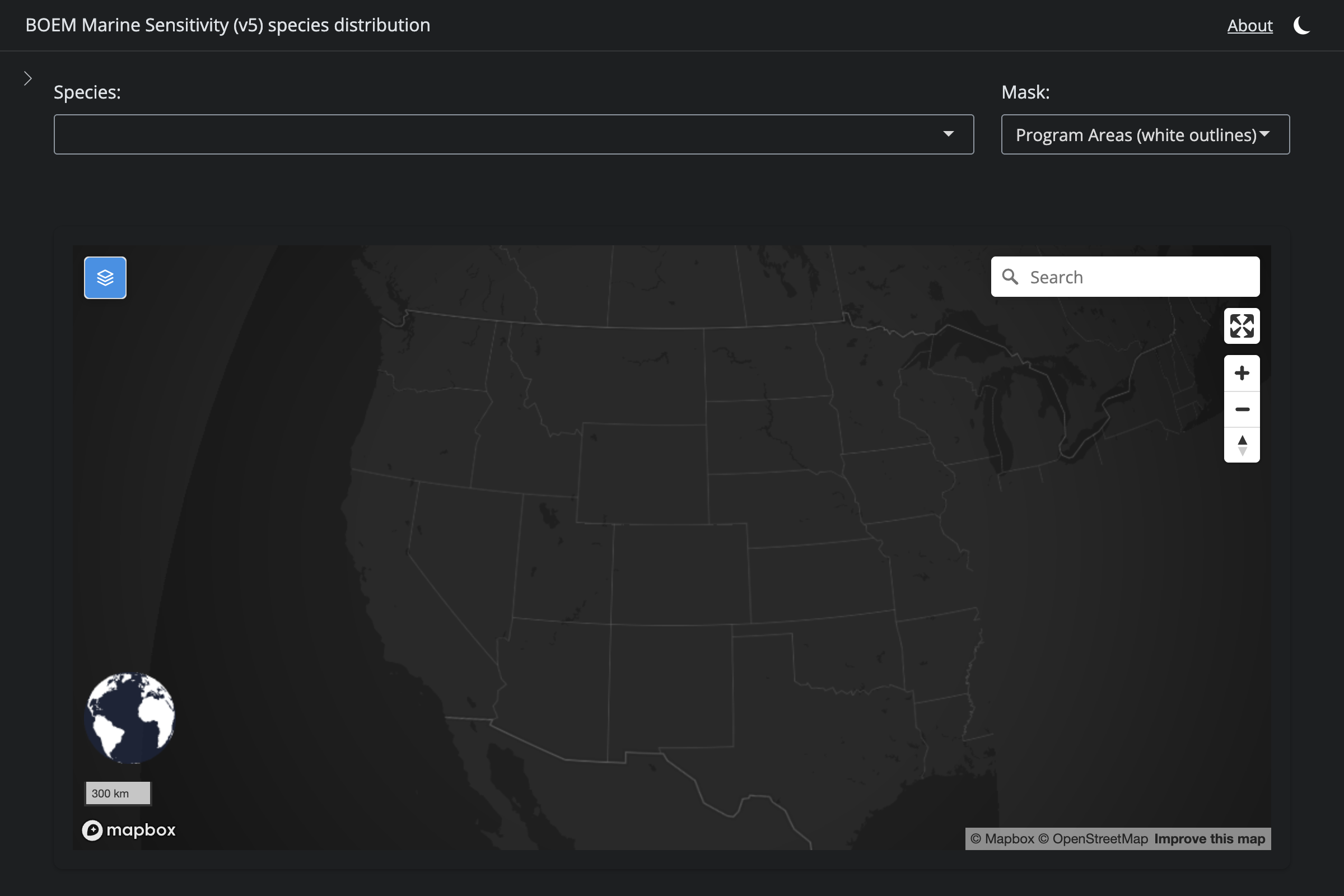
if (!is.na(sp$v6_mdl_seq)) {
url_v6 <- glue("https://app.marinesensitivity.org/mapsp/?mdl_seq={sp$v6_mdl_seq}&splash=false")
cat(glue("**v6** ([app link]({url_v6}))\n\n"))
png_v6 <- glue("figs/explore_goa-marmam/{sp_slug}_v6.png")
if (file.exists(here(png_v6))) {
cat(glue("\n\n"))
} else {
cat("*Screenshot not yet captured.*\n\n")
}
} else {
cat("**v6**: no merged model (`is_ok=FALSE`)\n\n")
}
cat(":::\n:::\n\n")
# tab: goa cell counts ----
cat("#### GOA Comparison\n\n")
d_sp_goa <- d_goa_cmp |>
filter(sp_scientific == sp$scientific_name) |>
select(-sp_scientific, -common_name)
if (nrow(d_sp_goa) > 0) {
tbl_html <- knitr::kable(d_sp_goa, digits = 4, format = "html",
caption = "v2 vs v6 scores in GOA Program Areas")
cat(tbl_html)
cat("\n\n")
} else {
cat("*Species not present in any GOA program area in either version.*\n\n")
}
# tab: database ----
cat("#### Datasets (v6)\n\n")
d_sp_ds <- d_summary_spp |>
filter(scientific_name == sp$scientific_name) |>
select(-scientific_name, -common_name)
if (nrow(d_sp_ds) > 0) {
tbl_html <- knitr::kable(d_sp_ds, format = "html",
caption = "v6 datasets and cell counts")
cat(tbl_html)
cat("\n\n")
} else {
cat("*No models in v6 database.*\n\n")
}
cat(":::\n\n")
}
```
```{r}
#| label: cleanup
#| include: false
dbDisconnect(con_sdm)
dbDisconnect(con_sdm_v2)
```